EMD-32791
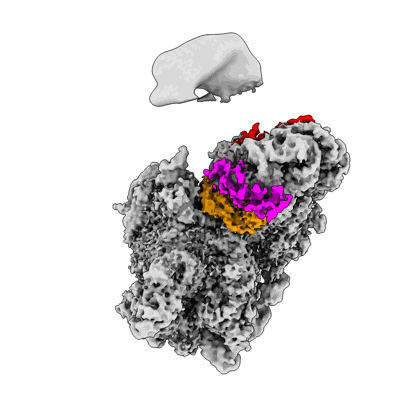
Cryo-EM structure of a yeast pre-40S ribosomal subunit - State Dis-E
EMD-32791
Single-particle3.5 Å
 Deposition: 05/02/2022
Deposition: 05/02/2022Map released: 19/10/2022
Last modified: 26/06/2024
Sample Organism:
Saccharomyces cerevisiae
Sample: Yeast pre-40S ribosomal subunit
Fitted models: 7wtm (Avg. Q-score: 0.451)
Deposition Authors: Cheng J, La Venuta G, Lau B ,
Berninghausen O,
Beckmann R,
Hurt E
,
Berninghausen O,
Beckmann R,
Hurt E  ,
Beckmann R
,
Beckmann R
Sample: Yeast pre-40S ribosomal subunit
Fitted models: 7wtm (Avg. Q-score: 0.451)
Deposition Authors: Cheng J, La Venuta G, Lau B
 ,
Berninghausen O,
Beckmann R,
Hurt E
,
Berninghausen O,
Beckmann R,
Hurt E  ,
Beckmann R
,
Beckmann R
In vitro structural maturation of an early stage pre-40S particle coupled with U3 snoRNA release and central pseudoknot formation.
Cheng J,
La Venuta G,
Lau B  ,
Berninghausen O,
Beckmann R,
Hurt E
,
Berninghausen O,
Beckmann R,
Hurt E 
(2022) Nucleic Acids Res , 50 , 11916 - 11923
 ,
Berninghausen O,
Beckmann R,
Hurt E
,
Berninghausen O,
Beckmann R,
Hurt E 
(2022) Nucleic Acids Res , 50 , 11916 - 11923
Abstract:
The transition of the 90S to the pre-40S pre-ribosome is a decisive step in eukaryotic small subunit biogenesis leading to a first pre-40S intermediate (state Dis-C or primordial pre-40S), where the U3 snoRNA keeps the nascent 18S rRNA locally immature. We in vitro reconstitute the ATP-dependent U3 release from this particle, catalyzed by the helicase Dhr1, and follow this process by cryo-EM revealing two successive pre-40S intermediates, Dis-D and Dis-E. The latter has lost not only U3 but all residual 90S factors including the GTPase Bms1. In vitro remodeling likewise induced the formation of the central pseudoknot, a universally conserved tertiary RNA structure that comprises the core of the small subunit decoding center. Thus, we could structurally reveal a key tertiary RNA folding step that is essential to form the active 40S subunit.
The transition of the 90S to the pre-40S pre-ribosome is a decisive step in eukaryotic small subunit biogenesis leading to a first pre-40S intermediate (state Dis-C or primordial pre-40S), where the U3 snoRNA keeps the nascent 18S rRNA locally immature. We in vitro reconstitute the ATP-dependent U3 release from this particle, catalyzed by the helicase Dhr1, and follow this process by cryo-EM revealing two successive pre-40S intermediates, Dis-D and Dis-E. The latter has lost not only U3 but all residual 90S factors including the GTPase Bms1. In vitro remodeling likewise induced the formation of the central pseudoknot, a universally conserved tertiary RNA structure that comprises the core of the small subunit decoding center. Thus, we could structurally reveal a key tertiary RNA folding step that is essential to form the active 40S subunit.
