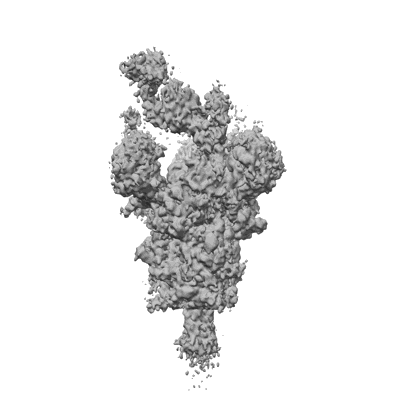
Cryo-EM structure of SARS-CoV-1 2p RBD in complex with W328-6H2(local refinement)
Resolution: 3.55 Å
EM Method: Single-particle
Fitted PDBs: 8jag
Q-score: 0.408
To Be Published
Cryo-EM structure of the Sars-Cov2 S trimer without RBDs
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8r87
Q-score: 0.496
Letscher H, Guilligay D, Sulbaran G, Amen A, Effantin G, Dereuddre-Bosquet N, Burger JA, Poniman M, Grobben M, Dergan Dylon S, Lemaitre J, Gros W, Galllouet A, Herate C, Relouzat F, Schoehn G, van der Werf S, van Gils MJ, Sanders RW, Poignard P, Le Grand R, Weissnhorn W
To Be Published
CryoEM structure of compound HNC-1664 bound with RdRP-RNA complex of SARS-CoV-2
Resolution: 3.29 Å
EM Method: Single-particle
Fitted PDBs: 8xko
Q-score: 0.486
Jia X, Jing X, Li M, Gao M, Zhong Y, Li E, Liu Y, Li R, Yao G, Liu Q, Zhou M, Hou Y, An L, Hong Y, Li S, Zhang J, Wang W, Zhang K, Gong P, Chiu S
Nat Commun (2024) 15 pp. 10750-10750 [ DOI: doi:10.1038/s41467-024-54918-3 Pubmed: 39737930 ]
- Rna (5'-r(p*up*up*cp*ap*up*gp*cp*up*ap*cp*gp*cp*gp*up*ap*g)-3') (5 kDa, RNA from synthetic construct)
- Cryoem structure of compound hnc-1644 bound with rdrp-rna (150 kDa, Complex from Severe acute respiratory syndrome coronavirus)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- [[(2~{r},3~{r},4~{s},5~{r})-4-fluoranyl-5-(5-iodanyl-4-methyl-pyrrolo[2,3-d]pyrimidin-7-yl)-3-oxidanyl-oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate (633 Da, Ligand)
- Rna (5'-r(p*cp*up*ap*cp*gp*cp*gp*up*ap*gp*cp*ap*up*g)-3') (4 kDa, RNA from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Non-structural protein 8 (23 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Rna-directed rna polymerase nsp12 (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with LLNL-199
Desautels TA, Arrildt KT, Zemla AT, Lau EY, Zhu F, Ricci D, Cronin S, Zost SJ, Binshtein E, Scheaffer SM, Dadonaite B, Petersen BK, Engdahl TB, Chen E, Handal LS, Hall L, Goforth JW, Vashchenko D, Nguyen S, Weilhammer DR, Lo JK, Rubinfeld B, Saada EA, Weisenberger T, Lee TH, Whitener B, Case JB, Ladd A, Silva MS, Haluska RM, Grzesiak EA, Earnhart CG, Hopkins S, Bates TW, Thackray LB, Segelke BW, Lillo AM, Sundaram S, Bloom JD, Diamond MS, Crowe Jr JE, Carnahan RH, Faissol DM
Nature (2024) 629 pp. 878-885 [ Pubmed: 38720086 DOI: doi:10.1038/s41586-024-07385-1 ]
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8ekd
Q-score: 0.433
Desautels TA, Arrildt KT, Zemla AT, Lau EY, Zhu F, Ricci D, Cronin S, Zost SJ, Binshtein E, Scheaffer SM, Dadonaite B, Petersen BK, Engdahl TB, Chen E, Handal LS, Hall L, Goforth JW, Vashchenko D, Nguyen S, Weilhammer DR, Lo JK, Rubinfeld B, Saada EA, Weisenberger T, Lee TH, Whitener B, Case JB, Ladd A, Silva MS, Haluska RM, Grzesiak EA, Earnhart CG, Hopkins S, Bates TW, Thackray LB, Segelke BW, Lillo AM, Sundaram S, Bloom JD, Diamond MS, Crowe Jr JE, Carnahan RH, Faissol DM
Nature (2024) 629 pp. 878-885 [ Pubmed: 38720086 DOI: doi:10.1038/s41586-024-07385-1 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Fab llnl-199 lc (12 kDa, Protein from Homo sapiens)
- Sars cov2 omicron ba.2 rbd complex with fab llnl-199 (Complex)
- Sars cov2 omicron ba.2 rbd (Complex from Severe acute respiratory syndrome coronavirus)
- Fab llnl-199 hc (14 kDa, Protein from Homo sapiens)
- Spike protein s2' (20 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Fab llnl-199 (Complex from Homo sapiens)
Aksu M, Kumar P, Guttler T, Taxer W, Gregor K, Mussil B, Rymarenko O, Stegmann KM, Dickmanns A, Gerber S, Reineking W, Schulz C, Henneck T, Mohamed A, Pohlmann G, Ramazanoglu M, Mese K, Gross U, Ben-Yedidia T, Ovadia O, Fischer DW, Kamensky M, Reichman A, Baumgartner W, von Kockritz-Blickwede M, Dobbelstein M, Gorlich D
Antiviral Res (2023) pp. 105778-105778 [ DOI: doi:10.1016/j.antiviral.2023.105778 Pubmed: 38065245 ]
- Vhh antibody (nanobody) ma16b06 (Protein from Vicugna pacos)
- Sars-cov-2 s (spike) protein (ba.1 variant) in complex with vhh ma16b06 (sub-volume of two adjacent rbd-vhh modules) (600 kDa, Complex from Severe acute respiratory syndrome coronavirus)
- Sars-cov-2 hexapro s (spike) glycoprotein (ba.1) (Protein from Severe acute respiratory syndrome coronavirus)
SARS-CoV-2 Omicron 1-RBD up Spike trimer complexed with three XG005 molecules
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 7ycy
Q-score: 0.26
To Be Published
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Xg005-vh (13 kDa, Protein from Homo sapiens)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Xg005-vl (11 kDa, Protein from Homo sapiens)
SARS-CoV-2 Omicron 2-RBD up Spike trimer complexed with three XG005 molecules
Resolution: 3.24 Å
EM Method: Single-particle
Fitted PDBs: 7ycz
Q-score: 0.352
To Be Published
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Xg005-vl (21 kDa, Protein from Homo sapiens)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Xg005-vh (23 kDa, Protein from Homo sapiens)
SARS-CoV-2 Omicron 1-RBD up spike trimer complexed with two XG005 Fab
Resolution: 3.58 Å
EM Method: Single-particle
Fitted PDBs: 7yd0
Q-score: 0.267
To Be Published
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Xg005-vl (21 kDa, Protein from Homo sapiens)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Xg005-vh (23 kDa, Protein from Homo sapiens)
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6A1 IgG