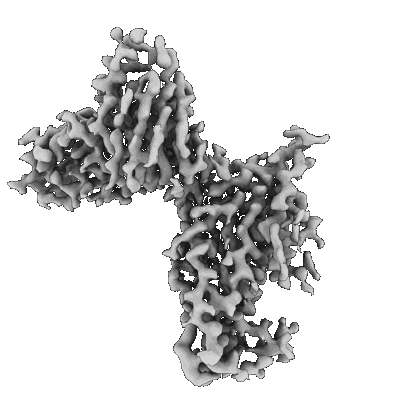
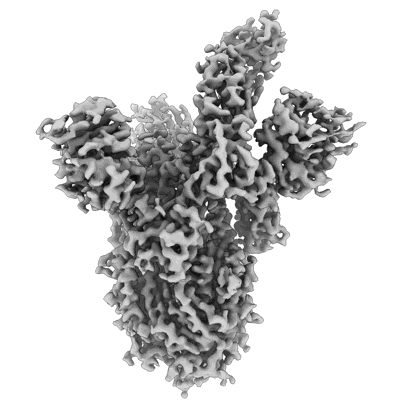
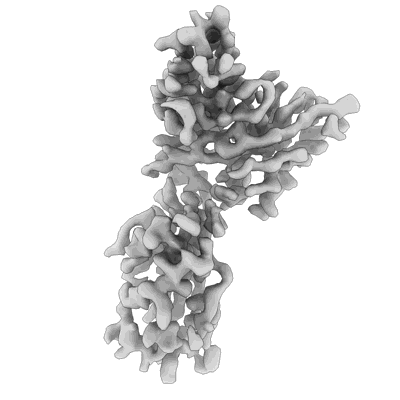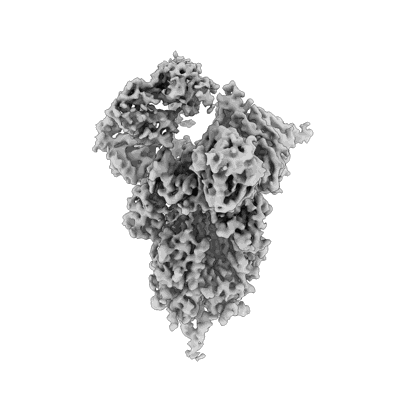Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8erq
Q-score: 0.493
Park YJ, Pinto D, Walls AC, Liu Z, De Marco A, Benigni F, Zatta F, Silacci-Fregni C, Bassi J, Sprouse KR, Addetia A, Bowen JE, Stewart C, Giurdanella M, Saliba C, Guarino B, Schmid MA, Franko NM, Logue JK, Dang HV, Hauser K, di Iulio J, Rivera W, Schnell G, Rajesh A, Zhou J, Farhat N, Kaiser H, Montiel-Ruiz M, Noack J, Lempp FA, Janer J, Abdelnabi R, Maes P, Ferrari P, Ceschi A, Giannini O, de Melo GD, Kergoat L, Bourhy H, Neyts J, Soriaga L, Purcell LA, Snell G, Whelan SPJ, Lanzavecchia A, Virgin HW, Piccoli L, Chu HY, Pizzuto MS, Corti D, Veesler D
Science (2022) 378 pp. 619-627 [ Pubmed: 36264829 DOI: doi:10.1126/science.adc9127 ]
- S2x324 fab heavy chain (13 kDa, Protein from Homo sapiens)
- S2x324 fab light chain (11 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 ba.1 spike ectodomain trimer in complex with the s2x324 neutralizing antibody fab fragment (local refinement of the rbd and s2x324) (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2x324 fab (Complex from Homo sapiens)
- Sars-cov-2 omicron ba.1 spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
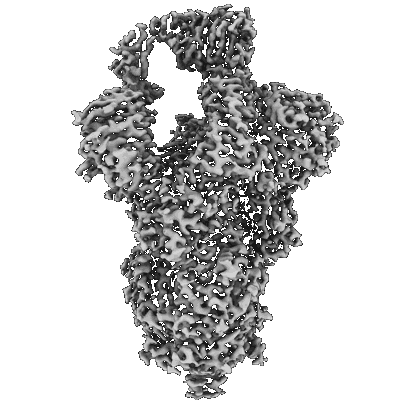
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8err
Q-score: 0.449
Park YJ, Pinto D, Walls AC, Liu Z, De Marco A, Benigni F, Zatta F, Silacci-Fregni C, Bassi J, Sprouse KR, Addetia A, Bowen JE, Stewart C, Giurdanella M, Saliba C, Guarino B, Schmid MA, Franko NM, Logue JK, Dang HV, Hauser K, di Iulio J, Rivera W, Schnell G, Rajesh A, Zhou J, Farhat N, Kaiser H, Montiel-Ruiz M, Noack J, Lempp FA, Janer J, Abdelnabi R, Maes P, Ferrari P, Ceschi A, Giannini O, de Melo GD, Kergoat L, Bourhy H, Neyts J, Soriaga L, Purcell LA, Snell G, Whelan SPJ, Lanzavecchia A, Virgin HW, Piccoli L, Chu HY, Pizzuto MS, Corti D, Veesler D
Science (2022) 378 pp. 619-627 [ Pubmed: 36264829 DOI: doi:10.1126/science.adc9127 ]
- Sars-cov-2 omicron ba.1 spike ectodomain trimer in complex with the s2x324 neutralizing antibody fab fragment (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron ba.1 spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2x324 fab (Complex from Homo sapiens)
- S2x324 light chain (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- S2x324 heavy chain (13 kDa, Protein from Homo sapiens)
Lempp FA, Soriaga LB, Montiel-Ruiz M, Benigni F, Noack J, Park YJ, Bianchi S, Walls AC, Bowen JE, Zhou J, Kaiser H, Joshi A, Agostini M, Meury M, Dellota Jr E, Jaconi S, Cameroni E, Martinez-Picado J, Vergara-Alert J, Izquierdo-Useros N, Virgin HW, Lanzavecchia A, Veesler D, Purcell LA, Telenti A, Corti D
Nature (2021) 598 pp. 342-347 [ DOI: doi:10.1038/s41586-021-03925-1 Pubmed: 34464958 ]
Lempp FA, Soriaga LB, Montiel-Ruiz M, Benigni F, Noack J, Park YJ, Bianchi S, Walls AC, Bowen JE, Zhou J, Kaiser H, Joshi A, Agostini M, Meury M, Dellota Jr E, Jaconi S, Cameroni E, Martinez-Picado J, Vergara-Alert J, Izquierdo-Useros N, Virgin HW, Lanzavecchia A, Veesler D, Purcell LA, Telenti A, Corti D
Nature (2021) 598 pp. 342-347 [ DOI: doi:10.1038/s41586-021-03925-1 Pubmed: 34464958 ]
SARS-CoV-2 spike protein bound to the S2P6 and S2M11 Fab fragments
Pinto D, Sauer MM, Czudnochowski N, Low JS, Tortorici MA, Housley MP, Noack J, Walls AC, Bowen JE, Guarino B, Rosen LE, di Iulio J, Jerak J, Kaiser H, Islam S, Jaconi S, Sprugasci N, Culap K, Abdelnabi R, Foo C, Coelmont L, Bartha I, Bianchi S, Silacci-Fregni C, Bassi J, Marzi R, Vetti E, Cassotta A, Ceschi A, Ferrari P, Cippa PE, Giannini O, Ceruti S, Garzoni C, Riva A, Benigni F, Cameroni E, Piccoli L, Pizzuto MS, Smithey M, Hong D, Telenti A, Lempp FA, Neyts J, Havenar-Daughton C, Lanzavecchia A, Sallusto F, Snell G, Virgin HW, Beltramello M, Corti D, Veesler D
Science (2021) 373 pp. 1109-1116 [ DOI: doi:10.1126/science.abj3321 Pubmed: 34344823 ]
- S2m11 fab fragment (Complex from Homo sapiens)
- Sars-cov-2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2p6 fab fragment (Complex from Homo sapiens)
- S2p6 and s2m11 fab fragments bound to the sars-cov-2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
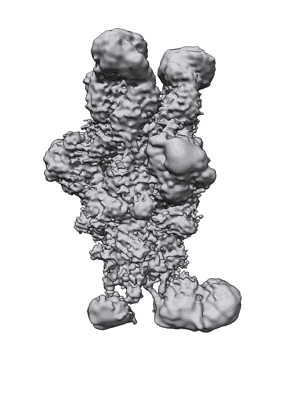
SARS-CoV-2 S glycoprotein in complex with S2X259 Fab
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7ra8
Q-score: 0.423
Tortorici MA, Czudnochowski N, Starr TN, Marzi R, Walls AC, Zatta F, Bowen JE, Jaconi S, Di Iulio J, Wang Z, De Marco A, Zepeda SK, Pinto D, Liu Z, Beltramello M, Bartha I, Housley MP, Lempp FA, Rosen LE, Dellota Jr E, Kaiser H, Montiel-Ruiz M, Zhou J, Addetia A, Guarino B, Culap K, Sprugasci N, Saliba C, Vetti E, Giacchetto-Sasselli I, Fregni CS, Abdelnabi R, Foo SC, Havenar-Daughton C, Schmid MA, Benigni F, Cameroni E, Neyts J, Telenti A, Virgin HW, Whelan SPJ, Snell G, Bloom JD, Corti D, Veesler D, Pizzuto MS
Nature (2021) 597 pp. 103-108 [ DOI: doi:10.1038/s41586-021-03817-4 Pubmed: 34280951 ]
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Severe acute respiratory syndrome coronavirus 2 (Complex)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 s glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Light chain fab s2x259 fab variable domain (11 kDa, Protein from Homo sapiens)
- S2x259 fab (Complex from Homo sapiens)
- Heavy chain fab s2x259 fab variable domain (13 kDa, Protein from Homo sapiens)
SARS-CoV-2 S bound to S2X259 Fab (local refinement of the RBD/S2X259 variable domains)
Tortorici MA, Czudnochowski N, Starr TN, Marzi R, Walls AC, Zatta F, Bowen JE, Jaconi S, Di Iulio J, Wang Z, De Marco A, Zepeda SK, Pinto D, Liu Z, Beltramello M, Bartha I, Housley MP, Lempp FA, Rosen LE, Dellota Jr E, Kaiser H, Montiel-Ruiz M, Zhou J, Addetia A, Guarino B, Culap K, Sprugasci N, Saliba C, Vetti E, Giacchetto-Sasselli I, Fregni CS, Abdelnabi R, Foo SC, Havenar-Daughton C, Schmid MA, Benigni F, Cameroni E, Neyts J, Telenti A, Virgin HW, Whelan SPJ, Snell G, Bloom JD, Corti D, Veesler D, Pizzuto MS
Nature (2021) 597 pp. 103-108 [ DOI: doi:10.1038/s41586-021-03817-4 Pubmed: 34280951 ]
- S2x259 fab light chain (11 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Severe acute respiratory syndrome coronavirus 2 (Complex)
- S2x259 fab heavy chain (13 kDa, Protein from Homo sapiens)
- Sars-cov-2 s (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2x259 fab (Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Resolution: 3.95 Å
EM Method: Single-particle
Fitted PDBs: 7r7n
Q-score: 0.411
Starr TN, Czudnochowski N, Liu Z, Zatta F, Park YJ, Addetia A, Pinto D, Beltramello M, Hernandez P, Greaney AJ, Marzi R, Glass WG, Zhang I, Dingens AS, Bowen JE, Tortorici MA, Walls AC, Wojcechowskyj JA, De Marco A, Rosen LE, Zhou J, Montiel-Ruiz M, Kaiser H, Dillen JR, Tucker H, Bassi J, Silacci-Fregni C, Housley MP, di Iulio J, Lombardo G, Agostini M, Sprugasci N, Culap K, Jaconi S, Meury M, Dellota Jr E, Abdelnabi R, Foo SC, Cameroni E, Stumpf S, Croll TI, Nix JC, Havenar-Daughton C, Piccoli L, Benigni F, Neyts J, Telenti A, Lempp FA, Pizzuto MS, Chodera JD, Hebner CM, Virgin HW, Whelan SPJ, Veesler D, Corti D, Bloom JD, Snell G
Nature (2021) 597 pp. 97-102 [ DOI: doi:10.1038/s41586-021-03807-6 Pubmed: 34261126 ]
- Sars-cov-2 spike in complex with the s2d106 neutralizing antibody fab fragment (Complex)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- S2d106 (Complex from Homo sapiens)
- Sars-cov-2 spike (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2d106 fab heavy chain (13 kDa, Protein from Homo sapiens)
- S2d106 fab light chain (11 kDa, Protein from Homo sapiens)
SARS-CoV-2 spike in complex with the S2D106 neutralizing antibody Fab fragment
Starr TN, Czudnochowski N, Liu Z, Zatta F, Park YJ, Addetia A, Pinto D, Beltramello M, Hernandez P, Greaney AJ, Marzi R, Glass WG, Zhang I, Dingens AS, Bowen JE, Tortorici MA, Walls AC, Wojcechowskyj JA, De Marco A, Rosen LE, Zhou J, Montiel-Ruiz M, Kaiser H, Dillen JR, Tucker H, Bassi J, Silacci-Fregni C, Housley MP, di Iulio J, Lombardo G, Agostini M, Sprugasci N, Culap K, Jaconi S, Meury M, Dellota Jr E, Abdelnabi R, Foo SC, Cameroni E, Stumpf S, Croll TI, Nix JC, Havenar-Daughton C, Piccoli L, Benigni F, Neyts J, Telenti A, Lempp FA, Pizzuto MS, Chodera JD, Hebner CM, Virgin HW, Whelan SPJ, Veesler D, Corti D, Bloom JD, Snell G
Nature (2021) 597 pp. 97-102 [ DOI: doi:10.1038/s41586-021-03807-6 Pubmed: 34261126 ]
Starr TN, Czudnochowski N, Liu Z, Zatta F, Park YJ, Addetia A, Pinto D, Beltramello M, Hernandez P, Greaney AJ, Marzi R, Glass WG, Zhang I, Dingens AS, Bowen JE, Tortorici MA, Walls AC, Wojcechowskyj JA, De Marco A, Rosen LE, Zhou J, Montiel-Ruiz M, Kaiser H, Dillen JR, Tucker H, Bassi J, Silacci-Fregni C, Housley MP, di Iulio J, Lombardo G, Agostini M, Sprugasci N, Culap K, Jaconi S, Meury M, Dellota Jr E, Abdelnabi R, Foo SC, Cameroni E, Stumpf S, Croll TI, Nix JC, Havenar-Daughton C, Piccoli L, Benigni F, Neyts J, Telenti A, Lempp FA, Pizzuto MS, Chodera JD, Hebner CM, Virgin HW, Whelan SPJ, Veesler D, Corti D, Bloom JD, Snell G
Nature (2021) 597 pp. 97-102 [ DOI: doi:10.1038/s41586-021-03807-6 Pubmed: 34261126 ]