Banach BB, Cerutti G, Fahad AS, Shen CH, Oliveira De Souza M, Katsamba PS, Tsybovsky Y, Wang P, Nair MS, Huang Y, Francino-Urdániz IM, Steiner PJ, Gutiérrez-González M, Liu L, López Acevedo SN, Nazzari AF, Wolfe JR, Luo Y, Olia AS, Teng IT, Yu J, Zhou T, Reddem ER, Bimela J, Pan X, Madan B, Laflin AD, Nimrania R, Yuen KY, Whitehead TA, Ho DD, Kwong PD, Shapiro L, DeKosky BJ
Cell Rep (2021) 37 [ Pubmed: 34587480 DOI: 10.1016/j.celrep.2021.109771 ]
Image sets:
Banach BB, Cerutti G, Fahad AS, Shen CH, Oliveira De Souza M, Katsamba PS, Tsybovsky Y, Wang P, Nair MS, Huang Y, Francino-Urdániz IM, Steiner PJ, Gutiérrez-González M, Liu L, López Acevedo SN, Nazzari AF, Wolfe JR, Luo Y, Olia AS, Teng IT, Yu J, Zhou T, Reddem ER, Bimela J, Pan X, Madan B, Laflin AD, Nimrania R, Yuen KY, Whitehead TA, Ho DD, Kwong PD, Shapiro L, DeKosky BJ
Cell Rep (2021) 37 [ Pubmed: 34587480 DOI: 10.1016/j.celrep.2021.109771 ]
Image sets:
Cryo-EM structure of SARS-CoV-2 spike at pH 4.0
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schön A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879 [ DOI: 10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
Image sets:
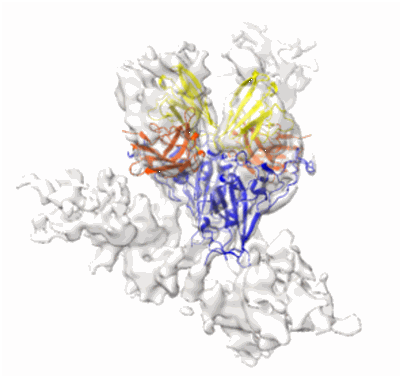
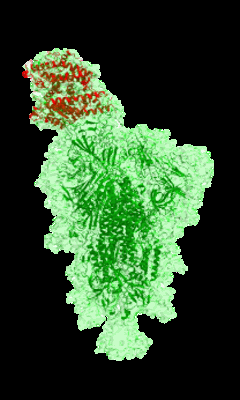
Cryo-EM structure of prefusion SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab
Resolution: 4.75 Å
EM Method: Single-particle
Fitted PDBs: 7ks9
Q-score: 0.223
Banach BB, Cerutti G, Fahad AS, Shen CH, Oliveira De Souza M, Katsamba PS, Tsybovsky Y, Wang P, Nair MS, Huang Y, Francino-Urdaniz IM, Steiner PJ, Gutierrez-Gonzalez M, Liu L, Lopez Acevedo SN, Nazzari AF, Wolfe JR, Luo Y, Olia AS, Teng IT, Yu J, Zhou T, Reddem ER, Bimela J, Pan X, Madan B, Laflin AD, Nimrania R, Yuen KY, Whitehead TA, Ho DD, Kwong PD, Shapiro L, DeKosky BJ
Cell Rep (2021) 37 pp. 109771-109771 [ Pubmed: 34587480 DOI: doi:10.1016/j.celrep.2021.109771 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Prefusion sars-cov-2 spike glycoprotein in complex with 910-30 fab (Complex from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 910-30 fab light chain (23 kDa, Protein from Homo sapiens)
- 910-30 fab heavy chain (23 kDa, Protein from Homo sapiens)
Cryo-EM map of SARS-CoV-2 spike glycoprotein in complex with 910-30 Fab (disrupted form)
Banach BB, Cerutti G, Fahad AS, Shen CH, Oliveira De Souza M, Katsamba PS, Tsybovsky Y, Wang P, Nair MS, Huang Y, Francino-Urdaniz IM, Steiner PJ, Gutierrez-Gonzalez M, Liu L, Lopez Acevedo SN, Nazzari AF, Wolfe JR, Luo Y, Olia AS, Teng IT, Yu J, Zhou T, Reddem ER, Bimela J, Pan X, Madan B, Laflin AD, Nimrania R, Yuen KY, Whitehead TA, Ho DD, Kwong PD, Shapiro L, DeKosky BJ
Cell Rep (2021) 37 pp. 109771-109771 [ Pubmed: 34587480 DOI: doi:10.1016/j.celrep.2021.109771 ]
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 5.5
Resolution: 3.85 Å
EM Method: Single-particle
Fitted PDBs: 7kne
Q-score: 0.253
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Single ace2-bound sars-cov-2 trimer spike at ph 5.5 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM Structure of Double ACE2-Bound SARS-CoV-2 Trimer Spike at pH 5.5
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 7knh
Q-score: 0.242
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Double ace2-bound sars-cov-2 trimer (Complex from Severe acute respiratory syndrome coronavirus 2)
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 5.5
Resolution: 3.91 Å
EM Method: Single-particle
Fitted PDBs: 7kni
Q-score: 0.144
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Triple ace2-bound sars-cov-2 trimer at ph 5.5 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Resolution: 3.39 Å
EM Method: Single-particle
Fitted PDBs: 7kmb
Q-score: 0.427
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- Ace2-rbd ph 7.4 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (140 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
Cryo-EM structure of triple ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Resolution: 3.64 Å
EM Method: Single-particle
Fitted PDBs: 7kms
Q-score: 0.262
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Triple ace2-bound sars-cov-2 trimer (Complex from Severe acute respiratory syndrome coronavirus 2)