EC 6.2.1.64: E1 NEDD8-activating enzyme
Reaction catalysed:
ATP + [NEDD8 protein] + [E1 NEDD8-activating enzyme]-L-cysteine = AMP + diphosphate + [E1 NEDD8-activating enzyme]-S-[NEDD8 protein]-yl-L-cysteine
Systematic name:
[NEDD8 protein]:[E1 NEDD8-activating enzyme] ligase (AMP-forming)
Alternative Name(s):
- NEDD-activating enzyme E1
GO terms
Biochemical function:
Biological process:
Cellular component:
Sequence families
 Protein families (Pfam)
Protein families (Pfam)
 InterPro annotations
InterPro annotations
IPR030468
Domain description:
NEDD8-activating enzyme E1 catalytic subunit, N-terminal domain
Occurring in:
Occurring in:
Structure domains
 CATH domains
CATH domains
3.40.50.720
Class:
Alpha Beta
Architecture:
3-Layer(aba) Sandwich
Topology:
Rossmann fold
Homology:
NAD(P)-binding Rossmann-like Domain
Occurring in:
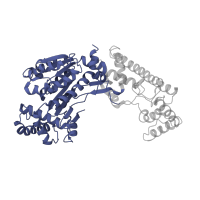
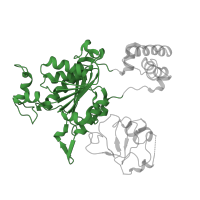
3.10.20.90
Class:
Alpha Beta
Architecture:
Roll
Topology:
Ubiquitin-like (UB roll)
Homology:
Phosphatidylinositol 3-kinase Catalytic Subunit; Chain A, domain 1
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1r4m</span> contains 4 copies of CATH domain <span class='highlight'>3.10.20.90 (Ubiquitin-like (UB roll))</span> in <span class='highlight'>Ubiquitin-like protein NEDD8</span>. Showing 1 copy in chain <span class='highlight'>C [auth I]</span>. The deposited structure of PDB entry 1r4m contains 4 copies of CATH domain 3.10.20.90 (Ubiquitin-like (UB roll)) in Ubiquitin-like protein NEDD8. Showing 1 copy in chain C [auth I].](https://www.ebi.ac.uk/pdbe/static/entry/1r4m_3_I_CATH_3.10.20.90_image-200x200.png)
3.10.20.260
Class:
Alpha Beta
Architecture:
Roll
Topology:
Ubiquitin-like (UB roll)
Homology:
NEDD8-activating enzyme E1, catalytic subunit
Occurring in:
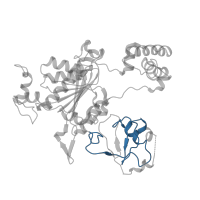





![The deposited structure of PDB entry <span class='highlight'>1r4m</span> contains 4 copies of Pfam domain <span class='highlight'>PF00240 (Ubiquitin family)</span> in <span class='highlight'>Ubiquitin-like protein NEDD8</span>. Showing 1 copy in chain <span class='highlight'>C [auth I]</span>. The deposited structure of PDB entry 1r4m contains 4 copies of Pfam domain PF00240 (Ubiquitin family) in Ubiquitin-like protein NEDD8. Showing 1 copy in chain C [auth I].](https://www.ebi.ac.uk/pdbe/static/entry/1r4m_3_I_Pfam_PF00240_image-200x200.png)




![The deposited structure of PDB entry <span class='highlight'>1r4m</span> contains 4 copies of SCOP domain <span class='highlight'>54237 (Ubiquitin-related)</span> in <span class='highlight'>Ubiquitin-like protein NEDD8</span>. Showing 1 copy in chain <span class='highlight'>C [auth I]</span>. The deposited structure of PDB entry 1r4m contains 4 copies of SCOP domain 54237 (Ubiquitin-related) in Ubiquitin-like protein NEDD8. Showing 1 copy in chain C [auth I].](https://www.ebi.ac.uk/pdbe/static/entry/1r4m_3_I_SCOP_54237_image-200x200.png)