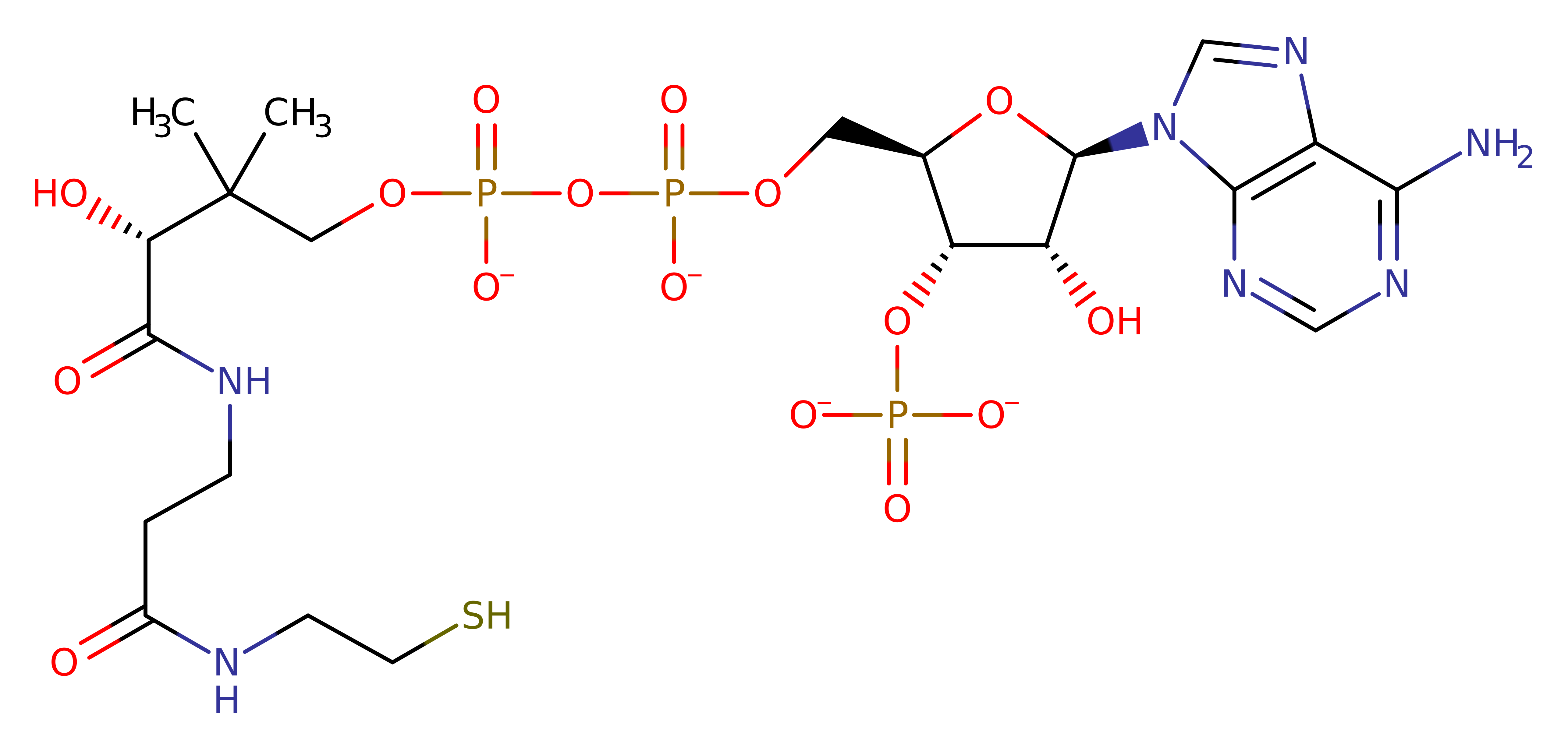
2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase
2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (also known as tetrahydrodipicolinate N-succinyltransferase or DapD) is part of the succinyl route of of lysine/diaminopimelic acid (DAP) biosynthesis. DAP biosynthesis is one of the two lysine biosynthesis pathways that are known to have evolved.
Reference Protein and Structure
- Sequence
-
P56220
 (2.3.1.117)
(2.3.1.117)
 (Sequence Homologues)
(PDB Homologues)
(Sequence Homologues)
(PDB Homologues)
- Biological species
-
unidentified prokaryotic organism (Bacteria)

- PDB
-
2tdt
- COMPLEX OF TETRAHYDRODIPICOLINATE N-SUCCINYLTRANSFERASE WITH 2-AMINOPIMELATE AND COENZYME A
(2.0 Å)



- Catalytic CATH Domains
-
2.160.10.10
 (see all for 2tdt)
(see all for 2tdt)
Enzyme Reaction (EC:2.3.1.117)
Enzyme Mechanism
Introduction
The mechanism of this protein is still unresolved, with some debate as to the exact role of Asp141A. It catalyses the conversion of the cyclic tetrahydrodipicolinate (THDP) into the acyclic compound N-succinyl L-2-amino-6-oxopimelate using succinyl-CoA. It is known to be activated by divalent metal ions, but it is still unclear if these are catalytic or not. A mechanism has been postulated where in a first step of the reaction succinyl-CoA delivers the succinyl group to the active site residue Asp141, which would become succinylated by forming an anhydride species. In a second step, the succinyl group is transferred onto the substrate THDP. It is also unclear whether the cyclic substrate THDP is hydrolysed before or concomitant with the transfer of the succinyl group.
Catalytic Residues Roles
| UniProt | PDB* (2tdt) | ||
| Glu189 | Glu189A(AA) | Thought to act as a general acid/base. | proton shuttle (general acid/base) |
| Gly166 (main-N) | Gly166A(AA) (main-N) | Acts to stabilise the negatively charged intermediates and transition states formed during the course of the reaction. | electrostatic stabiliser |
| Asp141 | Asp141A(AB) | Studies have suggested that this residue may be involved in covalent catalysis [PMID:19394346]. It may also act as a general acid/base and abstract a proton from the nucleophilic 2-amino group of the acceptor. | covalent catalysis |
Chemical Components
References
- Schuldt L et al. (2009), J Mol Biol, 389, 863-879. The three-dimensional Structure of a mycobacterial DapD provides insights into DapD diversity and reveals unexpected particulars about the enzymatic mechanism. DOI:10.1016/j.jmb.2009.04.046. PMID:19394346.
- Schnell R et al. (2012), PLoS One, 7, e31133-. Tetrahydrodipicolinate N-succinyltransferase and dihydrodipicolinate synthase from Pseudomonas aeruginosa: structure analysis and gene deletion. DOI:10.1371/journal.pone.0031133. PMID:22359568.
- Beaman TW et al. (2002), Protein Sci, 11, 974-979. Acyl group specificity at the active site of tetrahydridipicolinate N-succinyltransferase. DOI:10.1110/ps.4310102. PMID:11910040.
- Beaman TW et al. (1998), Biochemistry, 37, 10363-10369. The Conformational Change and Active Site Structure of TetrahydrodipicolinateN-Succinyltransferase†,‡. DOI:10.1021/bi980759b. PMID:9671504.
Catalytic Residues Roles
| Residue | Roles |
|---|---|
| Glu189A(AA) | proton shuttle (general acid/base) |
| Gly166A(AA) (main-N) | electrostatic stabiliser |
| Asp141A(AB) | covalent catalysis |