Tyrosine phenol-lyase
Tyrosine phenol-lyases (EC:4.1.99.2) (beta-tyrosinase), are pyridoxal-phosphate enzymes belonging to the beta-eliminating lyase family. They catalyse the reversible cleavage of the Cβ–Cγ bond of L-Tyr to produce phenol, ammonium, and pyruvate.
Reference Protein and Structure
- Sequence
-
P31013
 (4.1.99.2)
(4.1.99.2)
 (Sequence Homologues)
(PDB Homologues)
(Sequence Homologues)
(PDB Homologues)
- Biological species
-
Citrobacter freundii (Bacteria)

- PDB
-
2tpl
- TYROSINE PHENOL-LYASE FROM CITROBACTER INTERMEDIUS COMPLEX WITH 3-(4'-HYDROXYPHENYL)PROPIONIC ACID, PYRIDOXAL-5'-PHOSPHATE AND CS+ ION
(2.5 Å)



- Catalytic CATH Domains
-
3.40.640.10
 3.90.1150.10
3.90.1150.10  (see all for 2tpl)
(see all for 2tpl)
- Cofactors
- Pyridoxal 5'-phosphate(2-) (1)
Enzyme Reaction (EC:4.1.99.2)
Enzyme Mechanism
- Summary
- Step 1
- Step 2
- Step 3
- Step 4
- Step 5
- Step 6
- Step 7
- Step 8
- Step 9
- Step 10
- Step 11
- Products
- All Steps
Introduction
The tyrosine substrate initiates the first transaldimination reaction to displace the Lys257. Lysine then abstracts a proton from the substrate with PLP acting as an electron sink. A second proton transfer occurs (with Tyr71B) which activates the phenol moiety for elimination, which occurs next. The resulting 2-aminoprop-2-enoate product is released by a second transaldimination. 2-aminoprop-2-enoate then undergoes tautomerisation and subsequent hydrolysis, most likely outside the active site.
Catalytic Residues Roles
| UniProt | PDB* (2tpl) | ||
| Phe448 | Phe448A | Introduces steric strain in the substrates, destabilising the ground state and stabilising the quinoid intermediates. | ground state destabiliser, electrostatic stabiliser |
| Tyr71 | Tyr71B | Acts as a general acid/base. | proton acceptor, proton donor |
| Lys257 | Llp257A | This residue is covalently attached to the PLP cofactor in the ground state of the enzyme. It acts as a general acid/base during the course of the reaction, and is responsible for the transaldimination that releases the final product from the cofactor to regenerate the active site. | covalently attached, nucleofuge, nucleophile, proton acceptor, proton donor, electron pair acceptor, electron pair donor |
| Phe123 | Phe123A | Ensures that when the PLP cofactor is no longer covalently attached to the enzyme it remains in the correct orientation for the reaction to occur. | steric role |
| Asp214 | Asp214A | Forms a hydrogen bond with the nitrogen of the pyrimidine ring of the PLP cofactor, stabilising the cofactor and allowing it to act as an electron sink. | electrostatic stabiliser |
| Thr124, Arg381 | Thr124A, Arg381A | Helps stabilise the reactive intermediates and transition states formed during the course of the reaction. | electrostatic stabiliser |
Chemical Components
bimolecular nucleophilic addition, proton transfer, overall reactant used, schiff base formed, unimolecular elimination by the conjugate base, enzyme-substrate complex cleavage, overall product formed, enzyme-substrate complex formation, native state of cofactor regenerated, native state of enzyme regenerated, tautomerisation (not keto-enol), reaction occurs outside the enzyme, intramolecular eliminationReferences
- Milić D et al. (2011), J Am Chem Soc, 133, 16468-16476. Crystallographic Snapshots of Tyrosine Phenol-lyase Show That Substrate Strain Plays a Role in C–C Bond Cleavage. DOI:10.1021/ja203361g. PMID:21899319.
- Milić D et al. (2008), J Biol Chem, 283, 29206-29214. Insights into the catalytic mechanism of tyrosine phenol-lyase from X-ray structures of quinonoid intermediates. DOI:10.1074/jbc.M802061200. PMID:18715865.
- Milić D et al. (2006), Biochemistry, 45, 7544-7552. Structures of apo- and holo-tyrosine phenol-lyase reveal a catalytically critical closed conformation and suggest a mechanism for activation by K+ ions. DOI:10.1021/bi0601858. PMID:16768450.
- Barbolina MV et al. (2000), Protein Eng, 13, 207-215. Citrobacter freundii tyrosine phenol-lyase: the role of asparagine 185 in modulating enzyme function through stabilization of a quinonoid intermediate. DOI:10.1093/protein/13.3.207. PMID:10775663.
- Sundararaju B et al. (1997), Biochemistry, 36, 6502-6510. The crystal structure of Citrobacter freundii tyrosine phenol-lyase complexed with 3-(4'-hydroxyphenyl)propionic acid, together with site-directed mutagenesis and kinetic analysis, demonstrates that arginine 381 is required for substrate specificity. DOI:10.1021/bi962917+. PMID:9174368.

Step 1. In the first step of the initial transaldimination reaction the substrate tyrosine initiates a nucleophilic attack on the PLP cofactor.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Phe123A | steric role |
| Thr124A | electrostatic stabiliser |
| Asp214A | electrostatic stabiliser |
| Arg381A | electrostatic stabiliser |
| Phe448A | ground state destabiliser |
| Llp257A | covalently attached, proton acceptor, electron pair acceptor |
Chemical Components
ingold: bimolecular nucleophilic addition, proton transfer, overall reactant used
Step 2. In the final step of the transaldimination Lys257 is eliminated from the PLP cofactor.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Thr124A | electrostatic stabiliser |
| Asp214A | electrostatic stabiliser |
| Arg381A | electrostatic stabiliser |
| Phe448A | electrostatic stabiliser |
| Phe123A | steric role |
| Llp257A | nucleofuge |
Chemical Components
schiff base formed, ingold: unimolecular elimination by the conjugate base, enzyme-substrate complex cleavage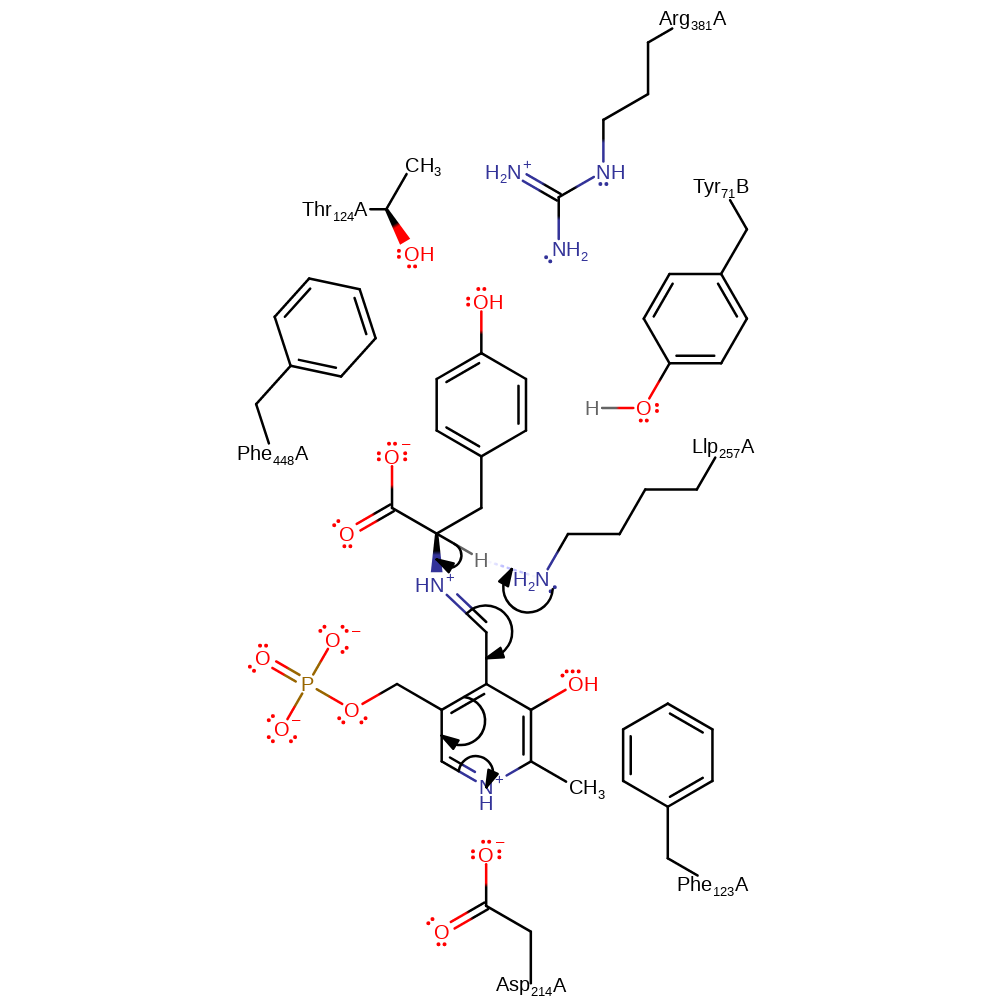
Step 3. Lys257 abstracts a proton from the covalently bound intermediate, which initiates a double bond rearrangement in which PLP acts as an electron sink.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Thr124A | electrostatic stabiliser |
| Asp214A | electrostatic stabiliser |
| Arg381A | electrostatic stabiliser |
| Phe448A | electrostatic stabiliser |
| Phe123A | steric role |
| Llp257A | proton acceptor |
Chemical Components
proton transfer
Step 4. A highly unstable quinoid species is formed by deprotonation of Tyr71B.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Thr124A | electrostatic stabiliser |
| Asp214A | electrostatic stabiliser |
| Arg381A | electrostatic stabiliser |
| Phe448A | electrostatic stabiliser |
| Phe123A | steric role |
| Tyr71B | proton donor |
Chemical Components
proton transfer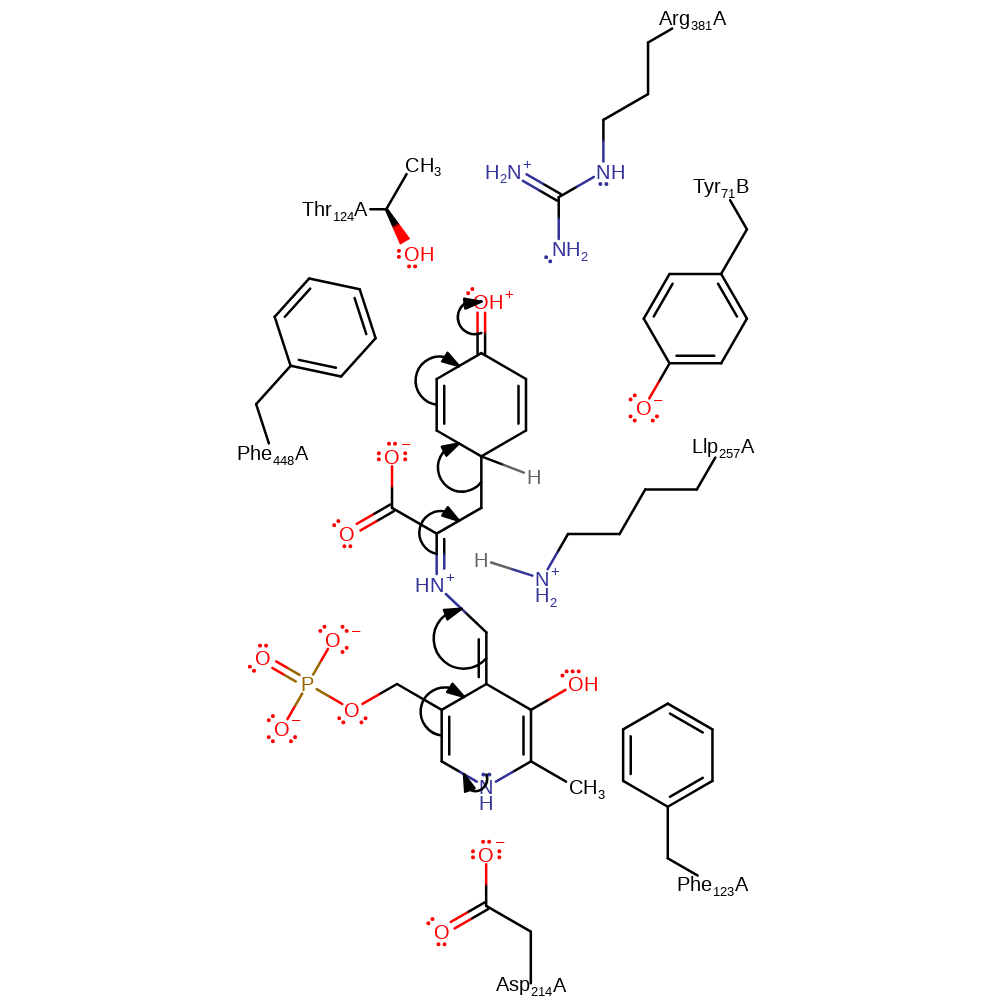
Step 5. The PLP cofactor initiates a double bond rearrangement that results in the elimination of the phenol group.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Thr124A | electrostatic stabiliser |
| Asp214A | electrostatic stabiliser |
| Arg381A | electrostatic stabiliser |
| Phe448A | electrostatic stabiliser |
| Phe123A | steric role |
Chemical Components
enzyme-substrate complex cleavage, ingold: unimolecular elimination by the conjugate base, overall product formed
Step 6. Tyr71B abstracts a proton from Lys257A, activating it for the reverse transaldimination reaction.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Phe123A | steric role |
| Asp214A | electrostatic stabiliser |
| Tyr71B | proton acceptor |
| Llp257A | proton donor |
Chemical Components
proton transfer
Step 7. Lys257A initiates a nucleophilic attack on the PLP cofactor in the first step of the final transaldimination reaction.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Phe123A | steric role |
| Asp214A | electrostatic stabiliser |
| Llp257A | proton donor, nucleophile |
Chemical Components
proton transfer, ingold: bimolecular nucleophilic addition, enzyme-substrate complex formationCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Asp214A | electrostatic stabiliser |
| Phe123A | steric role |
| Llp257A | electron pair donor |
Chemical Components
ingold: unimolecular elimination by the conjugate base, native state of cofactor regenerated, native state of enzyme regenerated, schiff base formed, enzyme-substrate complex cleavage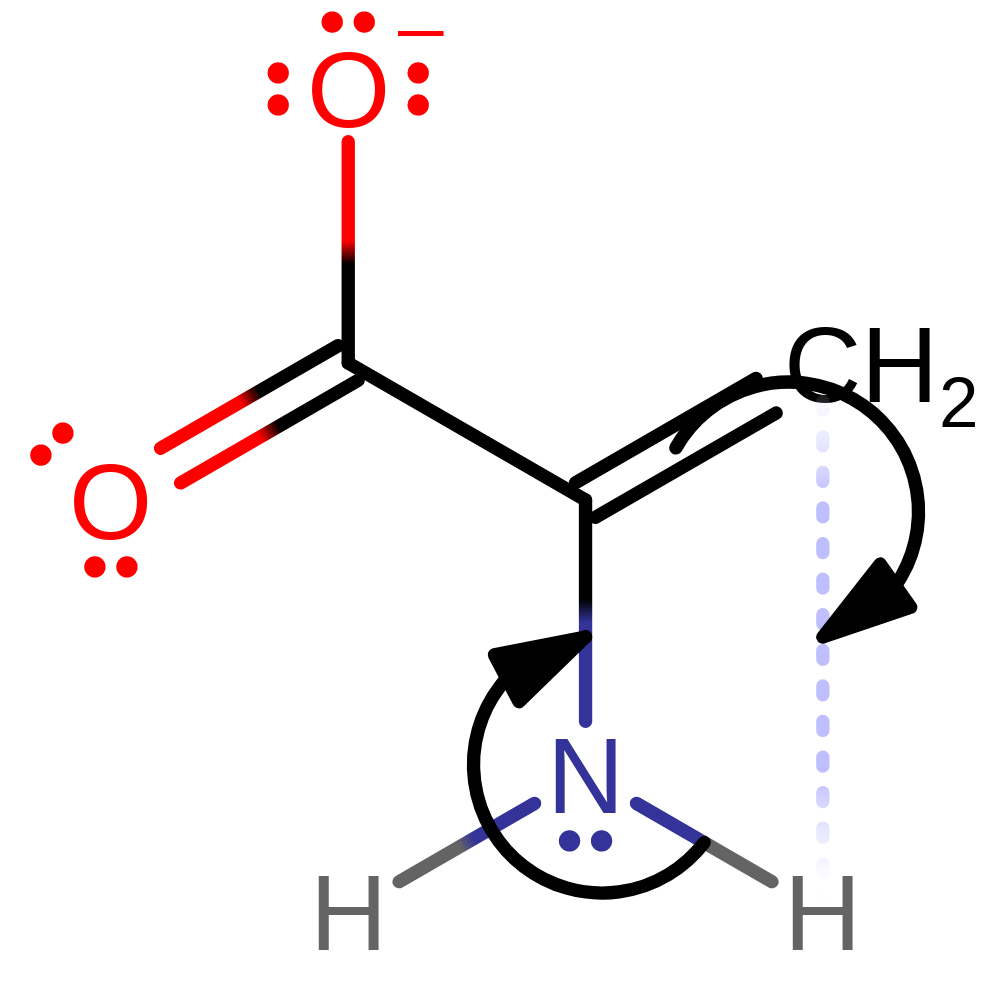
Step 9. It is likely that the rest of the mechanism occurs outside of the active site. The product undergoes tautomerisation.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|
Chemical Components
tautomerisation (not keto-enol), reaction occurs outside the enzymeCatalytic Residues Roles
| Residue | Roles |
|---|
Chemical Components
ingold: bimolecular nucleophilic addition, proton transfer, reaction occurs outside the enzyme
Step 11. The final products are formed by an intramolecular elimination.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|







 Download:
Download: