What is BioModels?
BioModels is an invaluable core resource for the systems biology/pharmacology community (4, 5, 6). It stores a vast collection of the literature-based mechanistic models in standard formats, many of which are physiologically and pharmaceutically relevant, and describe a wide range of biological/biomedical processes at different biological scales.
BioModels provides two sets of models:
- Models described in scientific literature (Individual / All models under quick search options)
- Models generated automatically from pathway resources (a branch of BioModels called Autogenerated Models)
We have manually validated the model components, as well as the structure and behaviour of a large proportion of literature-based models. This is done to ensure the models correspondence with the original publication. In Figure 2 you can see the general BioModels curation workflow.
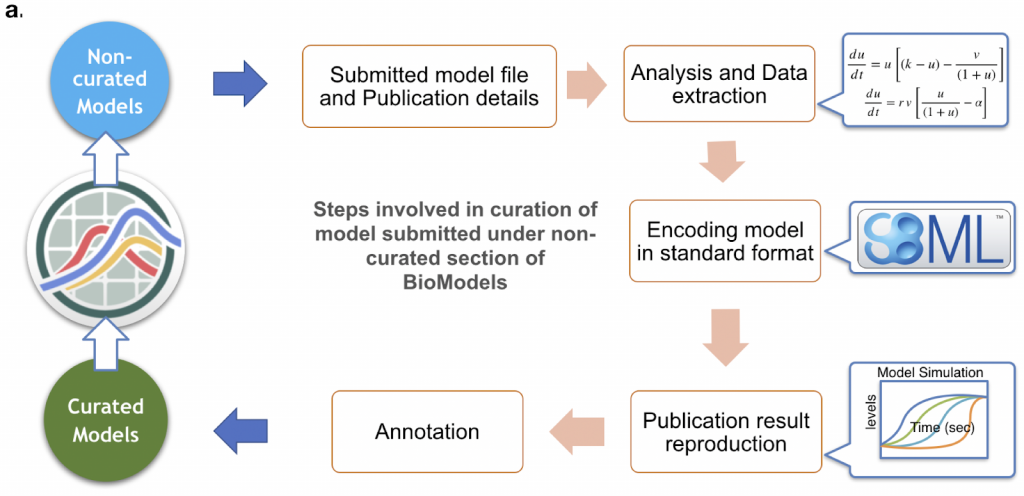
Figure 3 shows a specific example of the curation procedure for a mitogen-activated protein kinase cascades model.
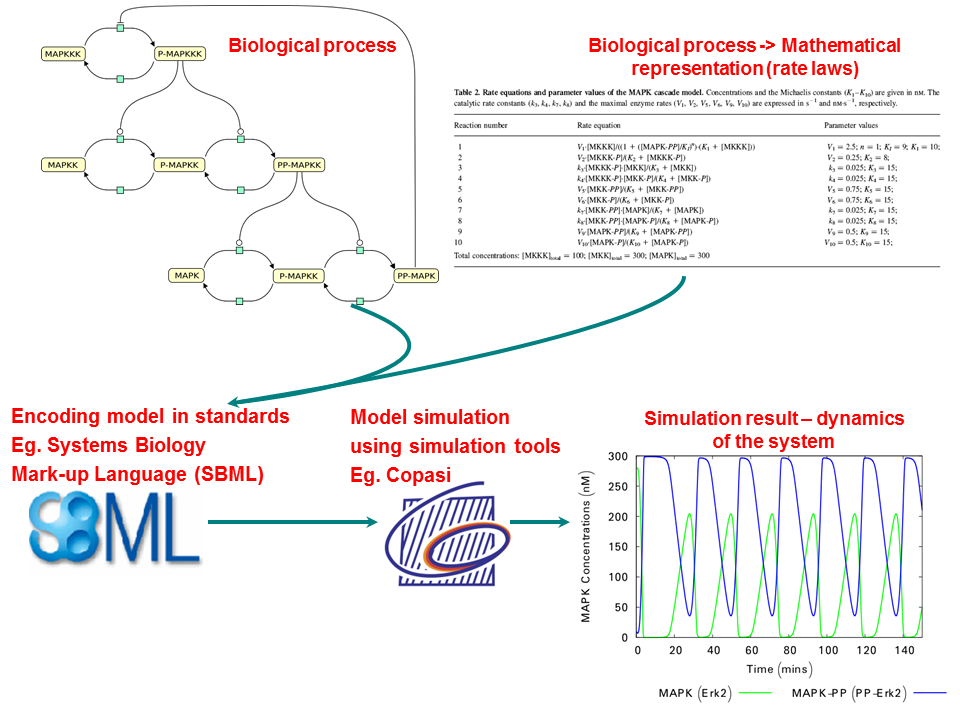
The model elements are also cross-linked to external database resources and ontologies, which enables you to precisely search models from the repository. BioModels is accessible through a web interface and programmatically through web services. Hosted models are freely available for use, modification and redistribution to all users under the terms of Creative Commons CC0.