Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
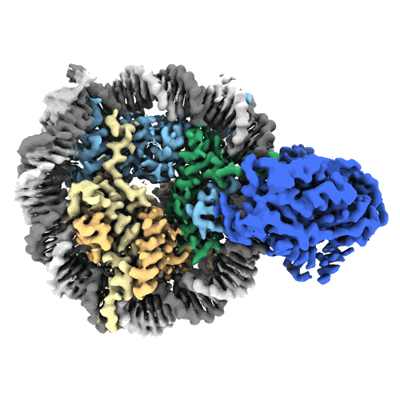
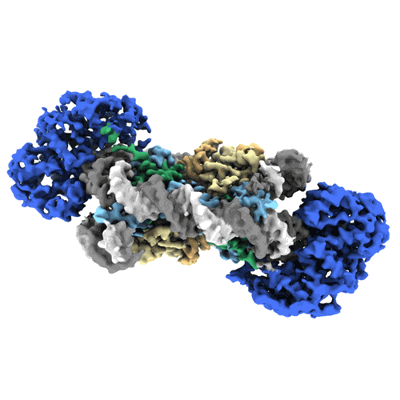
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ DOI: doi:10.1038/s41467-024-46178-y Pubmed: 38472177 ]
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna forward strand (DNA from synthetic construct)
- Histone h4 (Protein from Xenopus laevis)
- Histone h3.2 (Protein from Xenopus laevis)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna reverse strand (DNA from synthetic construct)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Histone h2b 1.1 (Protein from Xenopus laevis)
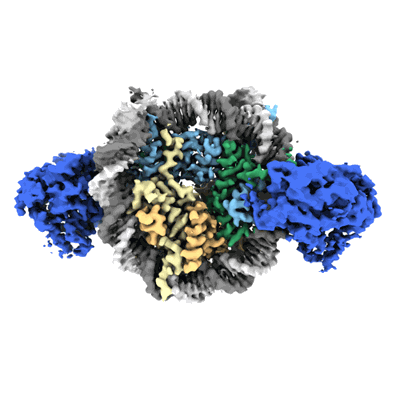
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8v6v
Q-score: 0.477
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ DOI: doi:10.1038/s41467-024-46178-y Pubmed: 38472177 ]
- Histone h3.2 (15 kDa, Protein from Xenopus laevis)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (122 kDa, Protein from Homo sapiens)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (64 kDa, DNA from synthetic construct)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Magnesium ion (24 Da, Ligand)
- Histone h2a type 1 (13 kDa, Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (63 kDa, DNA from synthetic construct)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8v7l
Q-score: 0.456
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ DOI: doi:10.1038/s41467-024-46178-y Pubmed: 38472177 ]
- Histone h3.2 (15 kDa, Protein from Xenopus laevis)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (122 kDa, Protein from Homo sapiens)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna (reverse strand) (64 kDa, DNA from synthetic construct)
- Histone h2a type 1 (13 kDa, Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Magnesium ion (24 Da, Ligand)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (63 kDa, DNA from synthetic construct)
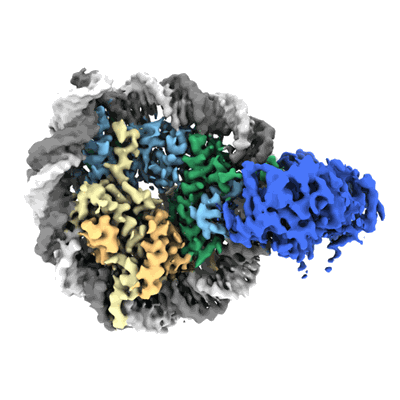
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ DOI: doi:10.1038/s41467-024-46178-y Pubmed: 38472177 ]
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Histone h4 (Protein from Xenopus laevis)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (DNA from synthetic construct)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (DNA from synthetic construct)
- Histone h3.2 (Protein from Xenopus laevis)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Histone h2b 1.1 (Protein from Xenopus laevis)
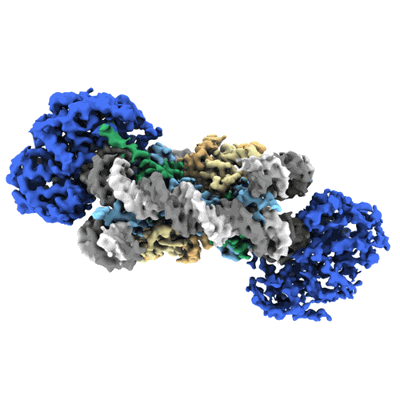
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ DOI: doi:10.1038/s41467-024-46178-y Pubmed: 38472177 ]
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Histone h4 (Protein from Xenopus laevis)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (DNA from synthetic construct)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (DNA from synthetic construct)
- Histone h3.2 (Protein from Xenopus laevis)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Histone h2b 1.1 (Protein from Xenopus laevis)
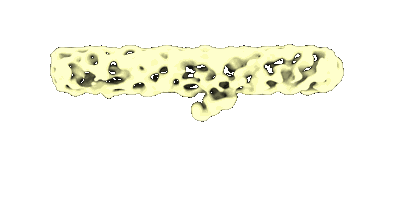
Type IV pilus from Pseudomonas PAO1 strain
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8tum
Q-score: 0.448
Thongchol J, Yu Z, Harb L, Lin Y, Koch M, Theodore M, Narsaria U, Shaevitz J, Gitai Z, Wu Y, Zhang J, Zeng L
Science (2024) 384 pp. eadl0635-eadl0635 [ DOI: doi:10.1126/science.adl0635 Pubmed: 38574145 ]
- Type iv major pilin protein pila (14 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Type iv twitching pilus (240 kDa, Cellular component from Pseudomonas aeruginosa)
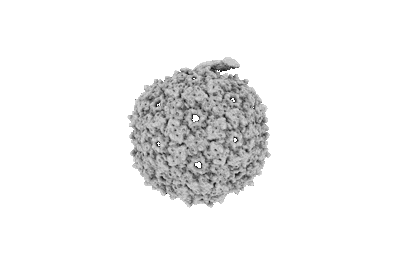
Asymmetric reconstruction of mature PP7 virions
Thongchol J, Yu Z, Harb L, Lin Y, Koch M, Theodore M, Narsaria U, Shaevitz J, Gitai Z, Wu Y, Zhang J, Zeng L
Science (2024) 384 pp. eadl0635-eadl0635 [ DOI: doi:10.1126/science.adl0635 Pubmed: 38574145 ]
- Type iv twitching pilus (240 kDa, Cellular component from Pseudomonas aeruginosa)
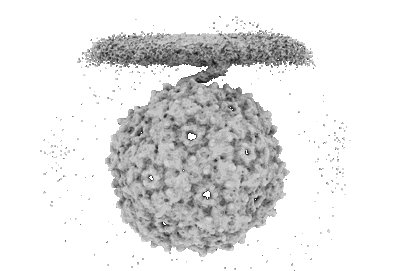
Type IV pilus from Pseudomonas PAO1 strain with PP7 Maturation protein
Resolution: 7.9 Å
EM Method: Single-particle
Fitted PDBs: 8tuw
Q-score: 0.087
Thongchol J, Yu Z, Harb L, Lin Y, Koch M, Theodore M, Narsaria U, Shaevitz J, Gitai Z, Wu Y, Zhang J, Zeng L
Science (2024) 384 pp. eadl0635-eadl0635 [ DOI: doi:10.1126/science.adl0635 Pubmed: 38574145 ]
- Type iv major pilin protein pila (14 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Maturation protein a (50 kDa, Protein from Pseudomonas phage PP7)
- Type iv twitching pilus (240 kDa, Cellular component from Pseudomonas aeruginosa)
The original consensus map of PP7 binding to T4P from PAO1
Thongchol J, Yu Z, Harb L, Lin Y, Koch M, Theodore M, Narsaria U, Shaevitz J, Gitai Z, Wu Y, Zhang J, Zeng L
Science (2024) 384 pp. eadl0635-eadl0635 [ DOI: doi:10.1126/science.adl0635 Pubmed: 38574145 ]
- Type iv twitching pilus (240 kDa, Cellular component from Pseudomonas aeruginosa)
Composite map of PP7 binding to type IV pilus from PAO1
Thongchol J, Yu Z, Harb L, Lin Y, Koch M, Theodore M, Narsaria U, Shaevitz J, Gitai Z, Wu Y, Zhang J, Zeng L
Science (2024) 384 pp. eadl0635-eadl0635 [ DOI: doi:10.1126/science.adl0635 Pubmed: 38574145 ]
- Type iv twitching pilus (240 kDa, Cellular component from Pseudomonas aeruginosa)