EMD-35623
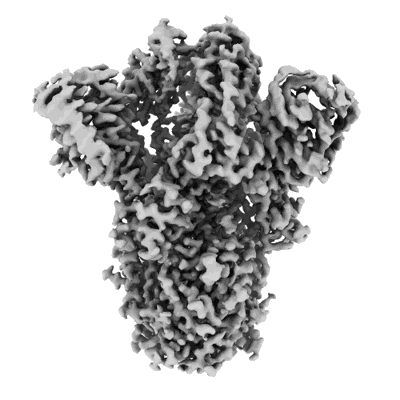
SARS-CoV-2 XBB.1 spike glycoprotein (closed-2 state)
EMD-35623
Single-particle2.51 Å
 Deposition: 13/03/2023
Deposition: 13/03/2023Map released: 24/05/2023
Last modified: 19/07/2023
Sample Organism:
Severe acute respiratory syndrome coronavirus 2
Sample: SARS-COV-2 XBB.1 spike glycoprotein
Fitted models: 8iot
Deposition Authors: Anraku Y ,
Kita S
,
Kita S  ,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Maenaka K
,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Maenaka K  ,
Hashiguchi T
,
Hashiguchi T 
Sample: SARS-COV-2 XBB.1 spike glycoprotein
Fitted models: 8iot
Deposition Authors: Anraku Y
 ,
Kita S
,
Kita S  ,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Maenaka K
,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Maenaka K  ,
Hashiguchi T
,
Hashiguchi T 
Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants.
Tamura T  ,
Ito J
,
Ito J  ,
Uriu K,
Zahradnik J
,
Uriu K,
Zahradnik J  ,
Kida I
,
Kida I  ,
Anraku Y
,
Anraku Y  ,
Nasser H
,
Nasser H  ,
Shofa M,
Oda Y,
Lytras S
,
Shofa M,
Oda Y,
Lytras S  ,
Nao N,
Itakura Y
,
Nao N,
Itakura Y  ,
Deguchi S,
Suzuki R,
Wang L
,
Deguchi S,
Suzuki R,
Wang L  ,
Begum MM,
Kita S
,
Begum MM,
Kita S  ,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Shimizu R,
Tsuda M
,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Shimizu R,
Tsuda M  ,
Kosugi Y,
Fujita S
,
Kosugi Y,
Fujita S  ,
Pan L,
Sauter D
,
Pan L,
Sauter D  ,
Yoshimatsu K
,
Yoshimatsu K  ,
Suzuki S
,
Suzuki S  ,
Asakura H,
Nagashima M,
Sadamasu K,
Yoshimura K,
Yamamoto Y,
Nagamoto T,
Schreiber G
,
Asakura H,
Nagashima M,
Sadamasu K,
Yoshimura K,
Yamamoto Y,
Nagamoto T,
Schreiber G  ,
Maenaka K
,
Maenaka K  ,
Hashiguchi T
,
Hashiguchi T  ,
Ikeda T
,
Ikeda T  ,
Fukuhara T
,
Fukuhara T  ,
Saito A
,
Saito A  ,
Tanaka S
,
Tanaka S  ,
Matsuno K
,
Matsuno K  ,
Takayama K
,
Takayama K  ,
Sato K
,
Sato K 
(2023) Nat Commun , 14 , 2800 - 2800
 ,
Ito J
,
Ito J  ,
Uriu K,
Zahradnik J
,
Uriu K,
Zahradnik J  ,
Kida I
,
Kida I  ,
Anraku Y
,
Anraku Y  ,
Nasser H
,
Nasser H  ,
Shofa M,
Oda Y,
Lytras S
,
Shofa M,
Oda Y,
Lytras S  ,
Nao N,
Itakura Y
,
Nao N,
Itakura Y  ,
Deguchi S,
Suzuki R,
Wang L
,
Deguchi S,
Suzuki R,
Wang L  ,
Begum MM,
Kita S
,
Begum MM,
Kita S  ,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Shimizu R,
Tsuda M
,
Yajima H,
Sasaki J,
Sasaki-Tabata K,
Shimizu R,
Tsuda M  ,
Kosugi Y,
Fujita S
,
Kosugi Y,
Fujita S  ,
Pan L,
Sauter D
,
Pan L,
Sauter D  ,
Yoshimatsu K
,
Yoshimatsu K  ,
Suzuki S
,
Suzuki S  ,
Asakura H,
Nagashima M,
Sadamasu K,
Yoshimura K,
Yamamoto Y,
Nagamoto T,
Schreiber G
,
Asakura H,
Nagashima M,
Sadamasu K,
Yoshimura K,
Yamamoto Y,
Nagamoto T,
Schreiber G  ,
Maenaka K
,
Maenaka K  ,
Hashiguchi T
,
Hashiguchi T  ,
Ikeda T
,
Ikeda T  ,
Fukuhara T
,
Fukuhara T  ,
Saito A
,
Saito A  ,
Tanaka S
,
Tanaka S  ,
Matsuno K
,
Matsuno K  ,
Takayama K
,
Takayama K  ,
Sato K
,
Sato K 
(2023) Nat Commun , 14 , 2800 - 2800
