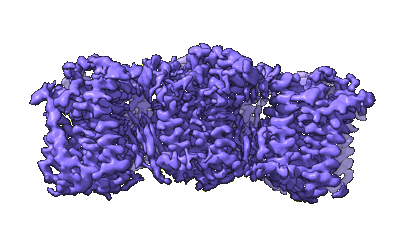
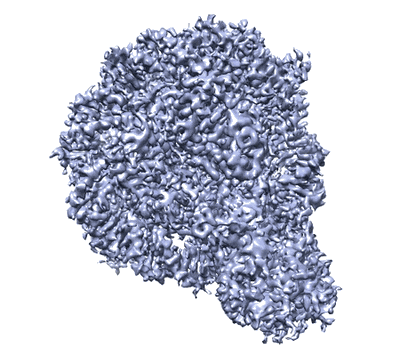
The cryo-EM structure of human sphingomyelin synthase-related protein in complex with ceramide
Resolution: 3.45 Å
EM Method: Single-particle
Fitted PDBs: 8ijq
Q-score: 0.461
Resolution: 3.29 Å
EM Method: Single-particle
Fitted PDBs: 8ijr
Q-score: 0.485
Hu K, Zhang Q, Chen Y, Yang J, Xia Y, Rao B, Li S, Shen Y, Cao M, Lu H, Qin A, Jiang XC, Yao D, Zhao J, Zhou L, Cao Y
Nat Struct Mol Biol (2024) 31 pp. 884-895 [ DOI: doi:10.1038/s41594-024-01237-2 Pubmed: 38388831 ]
- Sphingomyelin synthase-related protein 1 (48 kDa, Protein from Homo sapiens)
- (2s)-1-(hexadecanoyloxy)-3-hydroxypropan-2-yl (11z)-octadec-11-enoate (594 Da, Ligand)
- The cryo-em structure of human sphingomyelin synthase-related protein in complex with diacylglycerol/phosphoethanolamine (Complex from Homo sapiens)
- Phosphoric acid mono-(2-amino-ethyl) ester (141 Da, Ligand)
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 8w9w
Q-score: 0.377
Hu K, Zhang Q, Chen Y, Yang J, Xia Y, Rao B, Li S, Shen Y, Cao M, Lu H, Qin A, Jiang XC, Yao D, Zhao J, Zhou L, Cao Y
Nat Struct Mol Biol (2024) 31 pp. 884-895 [ DOI: doi:10.1038/s41594-024-01237-2 Pubmed: 38388831 ]
- ~{n}-[(~{z},2~{s},3~{r})-1,3-bis(oxidanyl)heptadec-4-en-2-yl]dodecanamide (467 Da, Ligand)
- Sphingomyelin synthase-related protein 1 (31 kDa, Protein from Homo sapiens)
- Phosphoric acid mono-(2-amino-ethyl) ester (141 Da, Ligand)
- The cryo-em structure of human sphingomyelin synthase-related protein in complex with ceramide/phosphoethanolamine (Complex from Homo sapiens)
Cryo-EM structure of human pyruvate carboxylase in apo state
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7wta
Q-score: 0.265
Chai P, Lan P, Li S, Yao D, Chang C, Cao M, Shen Y, Ge S, Wu J, Lei M, Fan X
Mol Cell (2022) 82 pp. 4116-4130.e6 [ DOI: doi:10.1016/j.molcel.2022.09.033 Pubmed: 36283412 ]
- Pyruvate carboxylase, mitochondrial (129 kDa, Protein from human)
- Pc (Complex from Homo sapiens)
- 5-(hexahydro-2-oxo-1h-thieno[3,4-d]imidazol-6-yl)pentanal (228 Da, Ligand)
Cryo-EM structure of human pyruvate carboxylase with acetyl-CoA
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 7wtb
Q-score: 0.313
Chai P, Lan P, Li S, Yao D, Chang C, Cao M, Shen Y, Ge S, Wu J, Lei M, Fan X
Mol Cell (2022) 82 pp. 4116-4130.e6 [ Pubmed: 36283412 DOI: doi:10.1016/j.molcel.2022.09.033 ]
- 5-(hexahydro-2-oxo-1h-thieno[3,4-d]imidazol-6-yl)pentanal (228 Da, Ligand)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Acetyl coenzyme *a (809 Da, Ligand)
- Pyruvate carboxylase, mitochondrial (129 kDa, Protein from human)
- Pc (Complex from Homo sapiens)
Cryo-EM structure of human pyruvate carboxylase with acetyl-CoA in the ground state
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 7wtc
Q-score: 0.216
Chai P, Lan P, Li S, Yao D, Chang C, Cao M, Shen Y, Ge S, Wu J, Lei M, Fan X
Mol Cell (2022) 82 pp. 4116-4130.e6 [ DOI: doi:10.1016/j.molcel.2022.09.033 Pubmed: 36283412 ]
- Pyruvate carboxylase, mitochondrial (129 kDa, Protein from human)
- Acetyl coenzyme *a (809 Da, Ligand)
- Pc (Complex from Homo sapiens)
- 5-(hexahydro-2-oxo-1h-thieno[3,4-d]imidazol-6-yl)pentanal (228 Da, Ligand)
Cryo-EM structure of human pyruvate carboxylase with acetyl-CoA in the intermediate state 1
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7wtd
Q-score: 0.226
Chai P, Lan P, Li S, Yao D, Chang C, Cao M, Shen Y, Ge S, Wu J, Lei M, Fan X
Mol Cell (2022) 82 pp. 4116-4130.e6 [ Pubmed: 36283412 DOI: doi:10.1016/j.molcel.2022.09.033 ]
- Pc (Complex from Homo sapiens)
- Coenzyme a (767 Da, Ligand)
- Pyruvate carboxylase, mitochondrial (129 kDa, Protein from human)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Cryo-EM structure of human pyruvate carboxylase with acetyl-CoA in the intermediate state 2
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7wte
Q-score: 0.36
Chai P, Lan P, Li S, Yao D, Chang C, Cao M, Shen Y, Ge S, Wu J, Lei M, Fan X
Mol Cell (2022) 82 pp. 4116-4130.e6 [ DOI: doi:10.1016/j.molcel.2022.09.033 Pubmed: 36283412 ]
- Pyruvate carboxylase, mitochondrial (129 kDa, Protein from human)
- Acetyl coenzyme *a (809 Da, Ligand)
- Pc (Complex from Homo sapiens)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Yeast Set2 bound to a nucleosome containing oncohistone mutations
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7ea5
Q-score: 0.473
Liu Y, Zhang Y, Xue H, Cao M, Bai G, Mu Z, Yao Y, Sun S, Fang D, Huang J
Cell Discov (2021) 7 pp. 32-32 [ Pubmed: 33972509 DOI: doi:10.1038/s41421-021-00261-6 ]
- Histone h4 (8 kDa, Protein from Xenopus laevis)
- S-adenosylmethionine (398 Da, Ligand)
- The set2-nucleosome(h3k36m)complex structure (Complex from Xenopus laevis)
- 601-dna (44 kDa, DNA from synthetic construct)
- Histone h2b (10 kDa, Protein from Xenopus laevis)
- Histone h2a (11 kDa, Protein from Xenopus laevis)
- Histone h3 (11 kDa, Protein from Xenopus laevis)
- Zinc ion (65 Da, Ligand)
- 601-dna (44 kDa, DNA from synthetic construct)
- Histone-lysine n-methyltransferase, h3 lysine-36 specific (26 kDa, Protein from Saccharomyces cerevisiae)
Human SETD2 bound to a nucleosome containing oncohistone mutations
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7ea8
Q-score: 0.509
Liu Y, Zhang Y, Xue H, Cao M, Bai G, Mu Z, Yao Y, Sun S, Fang D, Huang J
Cell Discov (2021) 7 pp. 32-32 [ DOI: doi:10.1038/s41421-021-00261-6 Pubmed: 33972509 ]
- Histone-lysine n-methyltransferase setd2 (28 kDa, Protein from Homo sapiens)
- Histone h4 (8 kDa, Protein from Homo sapiens)
- Histone-lysine n-methyltransferase setd2 (Complex from Homo sapiens)
- Human setd2-ncp(h3.3k36m)-sam complex structure (Complex from Homo sapiens)
- Histone h2b type 2-e (13 kDa, Protein from Homo sapiens)
- 601-dna (37 kDa, DNA from synthetic construct)
- Zinc ion (65 Da, Ligand)
- S-adenosylmethionine (398 Da, Ligand)
- Histone (Complex)
- 601-dna (37 kDa, DNA from synthetic construct)
- Histone h3.3 (11 kDa, Protein from Homo sapiens)
- Dna (Complex)
- Histone h2a type 1-d (11 kDa, Protein from Homo sapiens)