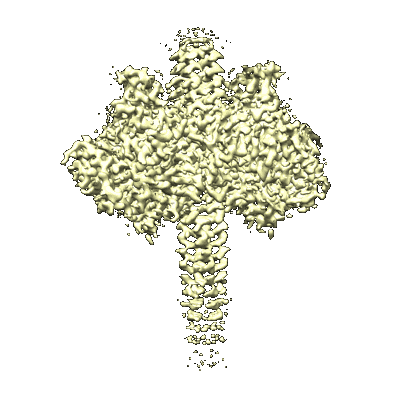
Langya Virus G glycoprotein (LayV G) with stabilizing mutations
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8tvi
Q-score: 0.552
Wang Z, McCallum M, Yan L, Gibson CA, Sharkey W, Park YJ, Dang HV, Amaya M, Person A, Broder CC, Veesler D
PNAS (2024) 121 pp. e2314990121-e2314990121 [ DOI: doi:10.1073/pnas.2314990121 Pubmed: 38593070 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Langya virus attachment glycoprotein g (68 kDa, Protein from Langya virus)
- Stabilized langya virus attachment glycoprotein layvg (Complex from Langya virus)
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8gwk
Q-score: 0.423
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- [(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methyl dihydrogen phosphate (371 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_rmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.31 Å
EM Method: Single-particle
Fitted PDBs: 8gw1
Q-score: 0.332
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_ump-nsp9 (350 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Manganese (ii) ion (54 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Uridine-5'-monophosphate (324 Da, Ligand)
- Water (18 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Replicase polyprotein 1ab (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna (25-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
lymphocytic choriomeningitis virus polymerase- Matrix Z Protein Complex (LCMV L-Z Complex)
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7x6v
Q-score: 0.393
Liu L, Wang P, Liu A, Zhang L, Yan L, Guo Y, Xiao G, Rao Z, Lou Z
Protein Cell (2023) 14 pp. 703-707 [ Pubmed: 37038286 DOI: doi:10.1093/procel/pwad018 ]
- Manganese (ii) ion (54 Da, Ligand)
- Ring finger protein z (10 kDa, Protein from Lymphocytic choriomeningitis virus (strain Armstrong))
- Lcmv polymerase l protein (Complex from Lymphocytic choriomeningitis virus (strain Armstrong))
- Matrix protein z (Complex from Lymphocytic choriomeningitis virus (strain Armstrong))
- The complex of lcmv polymerase with matrix protein z (Complex)
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (254 kDa, Protein from Lymphocytic choriomeningitis virus (strain Armstrong))
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 2.66 Å
EM Method: Single-particle
Fitted PDBs: 8gwe
Q-score: 0.019
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Replicase polyprotein 1a (8 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*ap*up*up*a)-3') (1 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_rna-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Helicase nsp13 (65 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.39 Å
EM Method: Single-particle
Fitted PDBs: 8gwf
Q-score: 0.29
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- E-rtc_rna-nsp9_gtp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (Complex)
- Nsp (Complex)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.37 Å
EM Method: Single-particle
Fitted PDBs: 8gwg
Q-score: 0.007
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Rna (Complex)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Protein from sars-cov-2 (Complex)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_smp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 8gwi
Q-score: 0.185
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (27-mer) (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_smp-nsp9_gtp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 8gwn
Q-score: 0.407
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_atmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 2.75 Å
EM Method: Single-particle
Fitted PDBs: 8gwb
Q-score: -0.011
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- E-rtc_rna-nsp9 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Manganese (ii) ion (54 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*ap*u)-3') (590 Da, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)