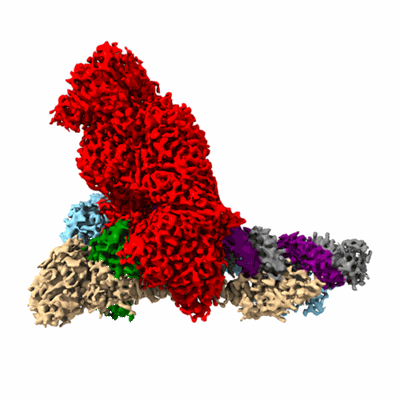
Cryo-EM structure of PEIP-Bs_enolase complex
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 7xml
Q-score: 0.468
Zhang K, Li S, Wang Y, Wang Z, Mulvenna N, Yang H, Zhang P, Chen H, Li Y, Wang H, Gao Y, Wigneshweraraj S, Matthews S, Zhang K, Liu B
Cell Rep (2022) 40 pp. 111026-111026 [ Pubmed: 35793626 DOI: doi:10.1016/j.celrep.2022.111026 ]
- Enolase (46 kDa, Protein from Bacillus subtilis (strain 168))
- Putative gene 60 protein (8 kDa, Protein from Bacillus phage SP01)
- Peip-bs_enolase complex (100 kDa, Complex)
- Bs_enolase (Complex from Bacillus subtilis (strain 168))
- Magnesium ion (24 Da, Ligand)
- Peip (Complex from Bacillus phage SP01)
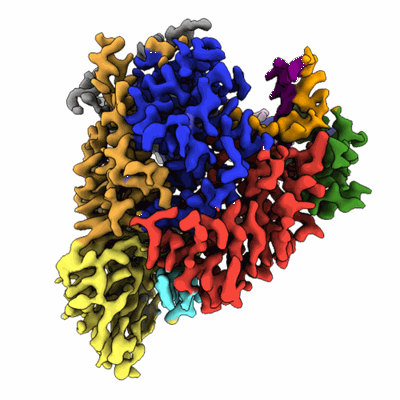
SARS-CoV-2 RdRP catalytic complex with T33-1 RNA
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7dte
Q-score: 0.528
Wu J, Wang H, Liu Q, Li R, Gao Y, Fang X, Zhong Y, Wang M, Wang Q, Rao Z, Gong P
Cell Rep (2021) 37 pp. 109882-109882 [ DOI: doi:10.1016/j.celrep.2021.109882 Pubmed: 34653416 ]
- Sars-cov-2 rna-dependent rna polymerase (Complex)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (33-mer) (10 kDa, RNA from Foot-and-mouth disease virus)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (Complex)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 rna-dependent rna polymerase catalytic complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna (57-mer) (18 kDa, RNA from Foot-and-mouth disease virus)
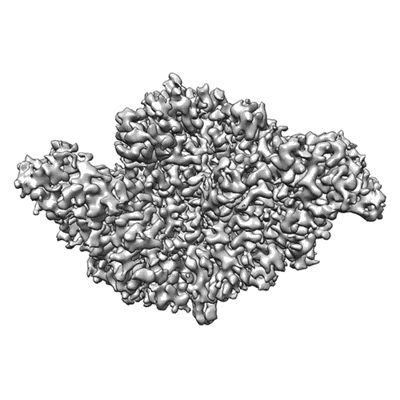
COVID-19 RNA-dependent RNA polymerase post-translocated catalytic complex
Resolution: 3.26 Å
EM Method: Single-particle
Fitted PDBs: 7bzf
Q-score: 0.47
Wang Q, Wu J, Wang H, Gao Y, Liu Q, Mu A, Ji W, Yan L, Zhu Y, Zhu C, Fang X, Yang X, Huang Y, Gao H, Liu F, Ge J, Sun Q, Yang X, Xu W, Liu Z, Yang H, Lou Z, Jiang B, Guddat LW, Gong P, Rao Z
Cell (2020) 182 pp. 417-428.e13 [ DOI: doi:10.1016/j.cell.2020.05.034 Pubmed: 32526208 ]
- Structure of covid-19 rna-dependent rna polymerase in complex with rna (155 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(*up*gp*up*up*cp*gp*ap*cp*gp*ap*cp*ap*cp*a)-3') (4 kDa, RNA from Foot-and-mouth disease virus)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (31-mer) (10 kDa, RNA from Foot-and-mouth disease virus)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
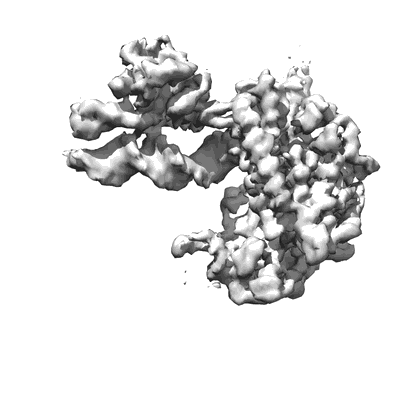
COVID-19 RNA-dependent RNA polymerase pre-translocated catalytic complex
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 7c2k
Q-score: 0.581
Wang Q, Wu J, Wang H, Gao Y, Liu Q, Mu A, Ji W, Yan L, Zhu Y, Zhu C, Fang X, Yang X, Huang Y, Gao H, Liu F, Ge J, Sun Q, Yang X, Xu W, Liu Z, Yang H, Lou Z, Jiang B, Guddat LW, Gong P, Rao Z
Cell (2020) 182 pp. 417-428.e13 [ Pubmed: 32526208 DOI: doi:10.1016/j.cell.2020.05.034 ]
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(*up*gp*up*up*cp*gp*ap*cp*gp*ap*cp*ap*cp*ap*gp*g*(f86)p*g)-3') (5 kDa, RNA from Foot-and-mouth disease virus)
- Covid-19 rna-dependent rna polymerase pre-translocated catalytic complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (29-mer) (9 kDa, RNA from Foot-and-mouth disease virus)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
Resolution: 4.5 Å
EM Method: Single-particle
Fitted PDBs: 6n7w
Q-score: 0.324
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Dna (25-mer) (7 kDa, DNA from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dna (77-mer) (23 kDa, DNA from Enterobacteria phage T7)
- Trxa (11 kDa, Protein from Escherichia coli)
- D5ae7a mutant gp5 dna polymerase complexed with trx, fork dna, and incoming dttp (from gp4-gp5 leading-strand complexes) (Complex from Enterobacteria phage T7)
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 6n9u
Q-score: 0.477
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Rna (5'-r(*ap*cp*cp*ap*g)-d(p*(doc))-3') (1 kDa, RNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Gp5 dna polymerase and two gp4 primase subunits complexed with trx, rna/dna hybrid, and incoming dttp (from gp4-gp5 lagging-strand complex lags1) (Complex from Enterobacteria phage T7)
- Dna (44-mer) (13 kDa, DNA from Enterobacteria phage T7)