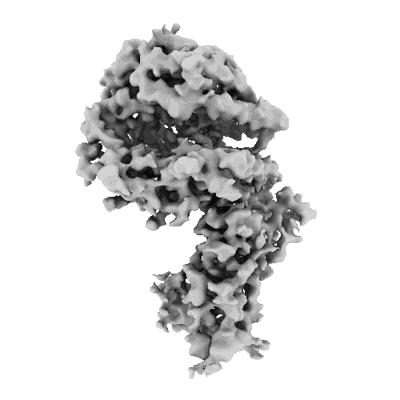
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2
Resolution: 3.78 Å
EM Method: Single-particle
Fitted PDBs: 8xln
Q-score: 0.447
Nomai T, Anraku Y, Kita S, Hashiguchi T, Maenaka K
bioRxiv (2024) [ DOI: doi:10.1101/2023.10.19.563209 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Processed angiotensin-converting enzyme 2 (70 kDa, Protein from Homo sapiens)
- Angiotensin-converting enzyme 2 (Complex from Homo sapiens)
- Spike glycoprotein (138 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 eg.5.1 spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 eg.5.1 spike glycoprotein in complex with ace2 (90 kDa, Complex)
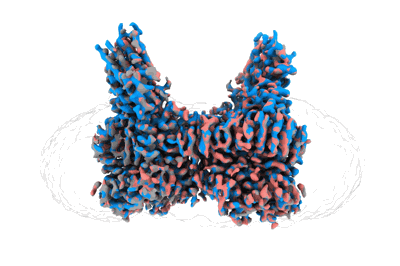
Structure of HamAB apo complex from the Escherichia coli Hachiman defense system
Resolution: 2.65 Å
EM Method: Single-particle
Fitted PDBs: 8vx9
Q-score: 0.418
Tuck OT, Adler BA, Armbruster EG, Lahiri A, Hu JJ, Zhou J, Pogliano J, Doudna JA
bioRxiv (2024) [ DOI: doi:10.1101/2024.02.29.582594 Pubmed: 38464307 ]
- Hamb (130 kDa, Protein from Escherichia coli)
- Apo hamab complex (160 kDa, Complex from Escherichia coli)
- Hama (28 kDa, Protein from Escherichia coli)
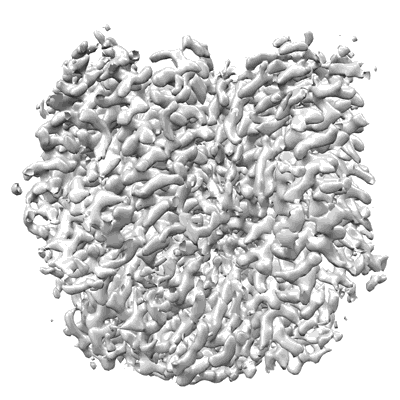
Structure of HamB-DNA complex, conformation 1, from the Escherichia coli Hachiman defense system
Resolution: 2.79 Å
EM Method: Single-particle
Fitted PDBs: 8vxa
Q-score: 0.462
Tuck OT, Adler BA, Armbruster EG, Lahiri A, Hu JJ, Zhou J, Pogliano J, Doudna JA
bioRxiv (2024) [ Pubmed: 38464307 DOI: doi:10.1101/2024.02.29.582594 ]
- Hamb (130 kDa, Protein from Escherichia coli)
- Hamb-dna complex (155 kDa, Complex from Escherichia coli)
- Dna (40-mer) (12 kDa, DNA from synthetic construct)
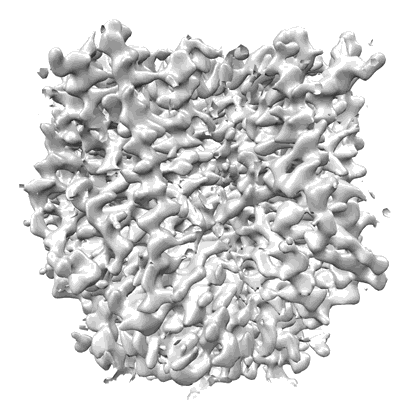
Structure of HamB-DNA complex, conformation 2, from the Escherichia coli Hachiman defense system
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 8vxc
Q-score: 0.408
Tuck OT, Adler BA, Armbruster EG, Lahiri A, Hu JJ, Zhou J, Pogliano J, Doudna JA
bioRxiv (2024) [ DOI: doi:10.1101/2024.02.29.582594 Pubmed: 38464307 ]
- Hamb-dna complex (155 kDa, Complex from Escherichia coli)
- Dna (40-mer) (12 kDa, DNA from synthetic construct)
- Hamb (130 kDa, Protein from Escherichia coli)
Resolution: 3.19 Å
EM Method: Single-particle
Fitted PDBs: 8vxy
Q-score: 0.315
Tuck OT, Adler BA, Armbruster EG, Lahiri A, Hu JJ, Zhou J, Pogliano J, Doudna JA
bioRxiv (2024) [ Pubmed: 38464307 DOI: doi:10.1101/2024.02.29.582594 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Plasmid dna (9 kDa, DNA from Escherichia coli)
- Hamb (130 kDa, Protein from Escherichia coli)
- Plasmid dna (9 kDa, DNA from Escherichia coli)
- Hama (28 kDa, Protein from Escherichia coli)
- Hama(e138a,k140a)b-plasmid dna complex (Complex from Escherichia coli)
Cryo-EM structure of HGSNAT-acetyl-CoA complex at pH 7.5
Resolution: 3.26 Å
EM Method: Single-particle
Fitted PDBs: 8tu9
Q-score: 0.508
Navratna V, Kumar A, Mosalaganti S
eLife (2024) 13 pp. RP93510- [ DOI: doi:10.7554/eLife.93510.1 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Acetyl coenzyme *a (809 Da, Ligand)
- Heparan acetyl-coa: alpha-glucosaminide n-acetyltransferase (hgsnat) (100 kDa, Complex from Homo sapiens)
- Enhanced green fluorescent protein,heparan-alpha-glucosaminide n-acetyltransferase,isoform 2 of heparan-alpha-glucosaminide n-acetyltransferase (100 kDa, Protein from Homo sapiens)
CryoEM structure of insect gustatory receptor BmGr9
Resolution: 2.85 Å
EM Method: Single-particle
Fitted PDBs: 8vc1
Q-score: 0.539
Frank HM, Walujkar S, Walsh RM, Laursen WJ, Theobald DL, Garrity PA, Gaudet R
bioRxiv (2023) [ Pubmed: 38187590 DOI: doi:10.1101/2023.12.19.572336 ]
- (3beta,14beta,17beta,25r)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en (544 Da, Ligand)
- Agonist-free bmgr9 homotetramer (218 kDa, Complex from Bombyx mori)
- (7s)-4-hydroxy-n,n,n-trimethyl-9-oxo-7-[(palmitoyloxy)methyl]-3,5,8-trioxa-4-phosphahexacosan-1-aminium 4-oxide (763 Da, Ligand)
- (7r,17e,20e)-4-hydroxy-n,n,n-trimethyl-9-oxo-7-[(palmitoyloxy)methyl]-3,5,8-trioxa-4-phosphahexacosa-17,20-dien-1-aminium 4-oxide (759 Da, Ligand)
- Gustatory receptor (54 kDa, Protein from Bombyx mori)
CryoEM structure of insect gustatory receptor BmGr9 in the presence of fructose
Resolution: 3.98 Å
EM Method: Single-particle
Fitted PDBs: 8vc2
Q-score: 0.376
Frank HM, Walujkar S, Walsh RM, Laursen WJ, Theobald DL, Garrity PA, Gaudet R
bioRxiv (2023) [ Pubmed: 38187590 DOI: doi:10.1101/2023.12.19.572336 ]
Structure of the mu opioid receptor bound to the antagonist nanobody NbE
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8qot
Q-score: 0.413
Yu J, Kumar A, Zhang X, Martin C, Raia P, Koehl A, Laeremans T, Steyaert J, Manglik A, Ballet S, Boland A, Stoeber M
bioRxiv (2023) [ Pubmed: 38106026 DOI: doi:10.1101/2023.12.06.570395 ]
- Nabfab hc (28 kDa, Protein from synthetic construct)
- Nabfab lc (25 kDa, Protein from synthetic construct)
- Mu-type opioid receptor (53 kDa, Protein from Mus musculus)
- Nanobody e (nbe) (18 kDa, Protein from Lama glama)
- Anti-fab nanobody (17 kDa, Protein from Lama glama)
- Mor-nbe-nabfab-antifab complex (120 kDa, Complex from Mus musculus)
PROTAC-mediated complex of KRAS with VHL/Elongin-B/Elongin-C/Cullin-2/Rbx1
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8qu8
Q-score: 0.306
Popow J, Farnaby W, Gollner A, Kofink C, Fischer G, Wurm M, Zollman D, Wijaya A, Mischerikow N, Hasenoehrl C, Prokofeva P, Arnhof H, Arce-Solano S, Bell S, Boeck G, Diers E, Frost A, Goodwin-Tindall J, Karolyi-Oezguer J, Khan S, Klawatsch T, Koegl M, Kousek R, Kratochvil B, Kropatsch K, Lauber A, McLennan R, Olt S, Peter D, Petermann O, Roessler V, Stolt-Bergner P, Strack P, Strauss E, Trainor N, Vetma V, Whitworth C, Zhong S, Quant J, Weinstabl H, Kuster B, Ettmayer P, Ciulli A
bioRxiv (2023) [ DOI: doi:10.1101/2023.10.24.563163 ]
- Cullin-2 (86 kDa, Protein from Homo sapiens)
- Kras/acbi3/vhl/elob/eloc/cul2/rbx1 (168 kDa, Complex from Homo sapiens)
- Gtpase kras (19 kDa, Protein from Homo sapiens)
- E3 ubiquitin-protein ligase rbx1, n-terminally processed (12 kDa, Protein from Homo sapiens)
- Von hippel-lindau disease tumor suppressor (24 kDa, Protein from Homo sapiens)
- Elongin-c (12 kDa, Protein from Homo sapiens)
- Guanosine-5'-diphosphate (443 Da, Ligand)
- (2s,4r)-1-[(2s)-2-[4-[4-[(3s)-4-[4-[5-[(4s)-2-azanyl-3-cyano-4-methyl-6,7-dihydro-5h-1-benzothiophen-4-yl]-1,2,4-oxadiazol-3-yl]pyrimidin-2-yl]-3-methyl-1,4-diazepan-1-yl]butoxy]-1,2,3-triazol-1-yl]-3-methyl-butanoyl]-n-[(1r)-1-[4-(4-methyl-1,3-thiazol-5-yl)phenyl]-2-oxidanyl-ethyl]-4-oxidanyl-pyrrolidine-2-carboxamide (1 kDa, Ligand)
- Zinc ion (65 Da, Ligand)
- Elongin-b (13 kDa, Protein from Homo sapiens)