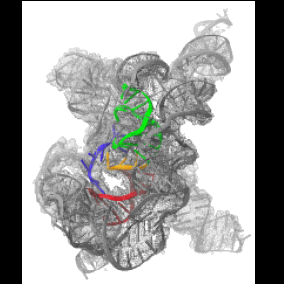
Cryo-EM structure of Tetrahymena ribozyme conformation 4 undergoing the second-step self-splicing
Resolution: 2.65 Å
EM Method: Single-particle
Fitted PDBs: 7ygc
Q-score: 0.454
Li S, Palo MZ, Zhang X, Pintilie G, Zhang K
Nat Commun (2023) 14 pp. 1294-1294 [ DOI: doi:10.1038/s41467-023-36724-5 Pubmed: 36928031 ]
- Rna (5'-r(*up*cp*g)-3') (911 Da, RNA from Tetrahymena thermophila)
- Magnesium ion (24 Da, Ligand)
- Rna (393-mer) (127 kDa, RNA from Tetrahymena thermophila)
- Rna (5'-r(*cp*cp*cp*up*cp*up*up*ap*ap*cp*c)-3') (3 kDa, RNA from Tetrahymena thermophila)
- Cryo-em structure of tetrahymena ribozyme conformation 4 undergoing the second-step self-splicing (130 kDa, Complex from Tetrahymena)
Misfolded Tetrahymena ribozyme conformation 1
Resolution: 3.53 Å
EM Method: Single-particle
Fitted PDBs: 7xsk
Q-score: 0.302
Li S, Palo MZ, Pintilie G, Zhang X, Su Z, Kappel K, Chiu W, Zhang K, Das R
PNAS (2022) 119 pp. e2209146119-e2209146119 [ DOI: doi:10.1073/pnas.2209146119 Pubmed: 36067294 ]
- Rna (388-mer) (125 kDa, RNA from Tetrahymena thermophila)
- Misfolded tetrahymena ribozyme conformation 1 (120 kDa, Complex from Tetrahymena thermophila)
Misfolded Tetrahymena ribozyme conformation 2
Resolution: 3.84 Å
EM Method: Single-particle
Fitted PDBs: 7xsl
Q-score: 0.279
Li S, Palo MZ, Pintilie G, Zhang X, Su Z, Kappel K, Chiu W, Zhang K, Das R
PNAS (2022) 119 pp. e2209146119-e2209146119 [ Pubmed: 36067294 DOI: doi:10.1073/pnas.2209146119 ]
- Rna (388-mer) (125 kDa, RNA from Tetrahymena thermophila)
- Misfolded tetrahymena ribozyme conformation 2 (120 kDa, Complex from Tetrahymena thermophila)
Misfolded Tetrahymena ribozyme conformation 3
Resolution: 4.01 Å
EM Method: Single-particle
Fitted PDBs: 7xsm
Q-score: 0.261
Li S, Palo MZ, Pintilie G, Zhang X, Su Z, Kappel K, Chiu W, Zhang K, Das R
PNAS (2022) 119 pp. e2209146119-e2209146119 [ DOI: doi:10.1073/pnas.2209146119 Pubmed: 36067294 ]
- Rna (388-mer) (125 kDa, RNA from Tetrahymena thermophila)
- Misfolded tetrahymena ribozyme conformation 3 (120 kDa, Complex from Tetrahymena thermophila)
Native Tetrahymena ribozyme conformation
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 7xsn
Q-score: 0.416
Li S, Palo MZ, Pintilie G, Zhang X, Su Z, Kappel K, Chiu W, Zhang K, Das R
PNAS (2022) 119 pp. e2209146119-e2209146119 [ DOI: doi:10.1073/pnas.2209146119 Pubmed: 36067294 ]
- Rna (387-mer) (125 kDa, RNA from Tetrahymena thermophila)
- Native tetrahymena ribozyme conformation (120 kDa, Complex from Tetrahymena thermophila)
Apo L-21 ScaI Tetrahymena ribozyme
Resolution: 3.14 Å
EM Method: Single-particle
Fitted PDBs: 7ez0
Q-score: 0.425
Holo L-16 ScaI Tetrahymena ribozyme
Resolution: 3.05 Å
EM Method: Single-particle
Fitted PDBs: 7ez2
Q-score: 0.485
Su Z, Zhang K, Kappel K, Li S, Palo MZ, Pintilie GD, Rangan R, Luo B, Wei Y, Das R, Chiu W
Nature (2021) 596 pp. 603-607 [ DOI: doi:10.1038/s41586-021-03803-w Pubmed: 34381213 ]
- Holo l-16 scai tetrahymena ribozyme s1 (2 kDa, RNA from Tetrahymena thermophila)
- Holo l-16 scai tetrahymena ribozyme s2 (1 kDa, RNA from Tetrahymena thermophila)
- Holo l-16 scai tetrahymena ribozyme (126 kDa, RNA from Tetrahymena thermophila)
- Holo l-16 scai tetrahymena ribozyme with substrates s1 and s2, and metal ions. (Complex from Tetrahymena thermophila)
- Magnesium ion (24 Da, Ligand)
Tetrahymena ribozyme models, 6.8 Angstrom resolution
Kappel K, Zhang K, Su Z, Watkins AM, Kladwang W, Li S, Pintilie G, Topkar VV, Rangan R, Zheludev IN, Yesselman JD, Chiu W, Das R
Nat Methods (2020) 17 pp. 699-707 [ Pubmed: 32616928 DOI: doi:10.1038/s41592-020-0878-9 ]
- Tetrahymena ribozyme (126 kDa, Complex from Tetrahymena thermophila)
- Rna (388-mer) (125 kDa, RNA from Tetrahymena thermophila)