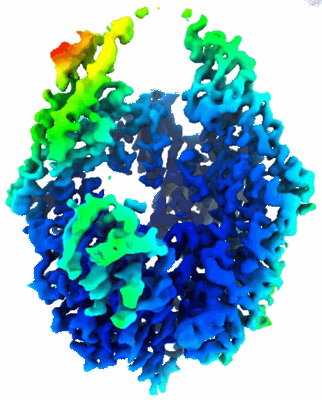
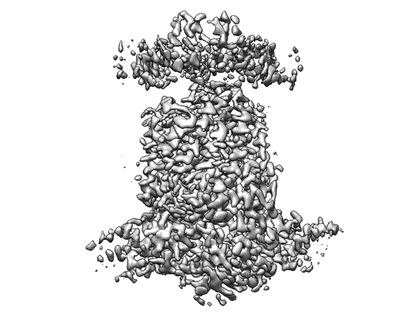
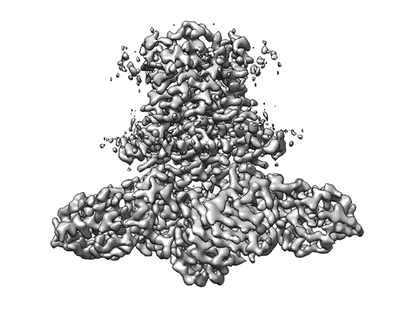
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 7z0s
Q-score: 0.64
Steinhilper R, Hoff G, Heider J, Murphy BJ
Nat Commun (2022) 13 pp. 5395-5395 [ DOI: doi:10.1038/s41467-022-32831-x Pubmed: 36104349 ]
- Iron/sulfur cluster (351 Da, Ligand)
- Phosphatidylethanolamine (734 Da, Ligand)
- Formate hydrogenlyase subunit 5 (66 kDa, Protein from Escherichia coli K-12)
- Formate hydrogenlyase subunit 6 (20 kDa, Protein from Escherichia coli K-12)
- Lauryl maltose neopentyl glycol (1 kDa, Ligand)
- Carbonmonoxide-(dicyano) iron (135 Da, Ligand)
- Formate hydrogenlyase subunit 2 (21 kDa, Protein from Escherichia coli K-12)
- Water (18 Da, Ligand)
- Cardiolipin (1 kDa, Ligand)
- Nickel (ii) ion (58 Da, Ligand)
- Formate hydrogenlyase subunit 3 (64 kDa, Protein from Escherichia coli K-12)
- 1-cis-9-octadecanoyl-2-cis-9-hexadecanoyl phosphatidyl glycerol (746 Da, Ligand)
- Escherichia coli formate hydrogenlyase complex without formate dehydrogenase h (Complex from Escherichia coli)
- Formate hydrogenlyase subunit 4 (33 kDa, Protein from Escherichia coli K-12)
- Formate hydrogenlyase subunit 7 (28 kDa, Protein from Escherichia coli K-12)
- Fe (iii) ion (55 Da, Ligand)
The structure of E. coli MutL bound to a 3' resected DNA end
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7p8v
Q-score: 0.448
Borsellini A, Lebbink JHG, Lamers MH
Nucleic Acids Res (2022) 50 pp. 6224-6234 [ DOI: doi:10.1093/nar/gkac432 Pubmed: 35670670 ]
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- 3' resected dna end (Complex from DNA molecule)
- Primer strand (3 kDa, DNA from DNA molecule)
- Dna mismatch repair protein mutl (68 kDa, Protein from Escherichia coli (strain K12))
- E. coli mutl n-terminal dimer bound to a 3' resected dna end (Complex)
- Magnesium ion (24 Da, Ligand)
- Template strand (6 kDa, DNA from DNA molecule)
- E. coli mutl n-terminal dimer (Complex from Escherichia coli K-12)
Structure of the autoinducer-2 exporter TqsA from E. coli
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7nb6
Q-score: 0.426
Khera R, Mehdipour AR, Bolla JR, Kahnt J, Welsch S, Ermler U, Muenke C, Robinson CV, Hummer G, Xie H, Michel H
EMBO J (2022) 41 pp. e109990-e109990 [ Pubmed: 35698912 DOI: doi:10.15252/embj.2021109990 ]
- Ai-2 transport protein tqsa (37 kDa, Protein from Escherichia coli (strain K12))
- Pentameric tqsa (187 kDa, Complex from Escherichia coli)
Structure of the AI-2 exporter family protein YdiK from E. coli
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7ot9
Q-score: 0.499
Khera R, Mehdipour AR, Bolla JR, Kahnt J, Welsch S, Ermler U, Muenke C, Robinson CV, Hummer G, Xie H, Michel H
EMBO J (2022) 41 pp. e109990-e109990 [ Pubmed: 35698912 DOI: doi:10.15252/embj.2021109990 ]
- Ai-2e member ydik (39 kDa, Protein from Escherichia coli (strain K12))
- Pentameric ydik (199 kDa, Complex from Escherichia coli (strain K12))
The structure of MutS bound to two molecules of AMPPNP
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7oto
Q-score: 0.499
Borsellini A, Kunetsky V, Friedhoff P, Lamers MH
Nat Struct Mol Biol (2022) 29 pp. 59-66 [ Pubmed: 35013597 DOI: doi:10.1038/s41594-021-00707-1 ]
- Dna mismatch repair protein muts (190 kDa, Cellular component from Escherichia coli)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Dna mismatch repair protein muts (90 kDa, Protein from Escherichia coli (strain K12))
- Magnesium ion (24 Da, Ligand)
The structure of MutS bound to two molecules of ADP-Vanadate
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 7ou0
Q-score: 0.413
Borsellini A, Kunetsky V, Friedhoff P, Lamers MH
Nat Struct Mol Biol (2022) 29 pp. 59-66 [ Pubmed: 35013597 DOI: doi:10.1038/s41594-021-00707-1 ]
- Magnesium ion (24 Da, Ligand)
- Dna mismatch repair protein muts (90 kDa, Protein from Escherichia coli (strain K12))
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Dna mismatch repair protein muts (190 kDa, Cellular component from Escherichia coli)
- Vanadate ion (114 Da, Ligand)
High resolution structure of cytochrome bd-II oxidase from E. coli
Resolution: 2.06 Å
EM Method: Single-particle
Fitted PDBs: 7oy2
Q-score: 0.7
Grund TN, Radloff M, Wu D, Goojani HG, Witte LF, Josting W, Buschmann S, Muller H, Elamri I, Welsch S, Schwalbe H, Michel H, Bald D, Safarian S
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2114013118 Pubmed: 34873041 ]
- Cytochrome bd-ii ubiquinol oxidase subunit 1 (57 kDa, Protein from Escherichia coli K-12)
- Cardiolipin (1 kDa, Ligand)
- Cytochrome bd-ii ubiquinol oxidase subunit 2 (42 kDa, Protein from Escherichia coli K-12)
- 2-[(2~{e},6~{e},10~{z},14~{e},18~{e},22~{e},26~{e})-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaenyl]naphthalene-1,4-dione (703 Da, Ligand)
- Heme b/c (618 Da, Ligand)
- Water (18 Da, Ligand)
- Cis-heme d hydroxychlorin gamma-spirolactone (632 Da, Ligand)
- Cytochrome bd-ii oxidase from e. coli composed of subunit appb, appc and appx (104 kDa, Complex from Escherichia coli (strain K12))
- (2s)-3-(hexadecanoyloxy)-2-[(9z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate (760 Da, Ligand)
- Oxygen molecule (31 Da, Ligand)
- Putative cytochrome bd-ii ubiquinol oxidase subunit appx (6 kDa, Protein from Escherichia coli K-12)
Cryo-EM structure of P.aeruginosa MlaFEBD with ADP-V
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7ch8
Q-score: 0.401
Zhou C, Shi H, Zhang M, Zhou L, Xiao L, Feng S, Im W, Zhou M, Zhang X, Huang Y
J Mol Biol (2021) 433 pp. 166986-166986 [ DOI: doi:10.1016/j.jmb.2021.166986 Pubmed: 33845086 ]
- E.coli mlafeb bond with amppnp (Complex from Escherichia coli K-12)
- Mlad domain-containing protein (16 kDa, Protein from Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1))
- Probable permease of abc transporter (28 kDa, Protein from Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1))
- Probable atp-binding component of abc transporter (29 kDa, Protein from Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1))
- Adp metavanadate (527 Da, Ligand)
- 2-(hexadecanoyloxy)-1-[(phosphonooxy)methyl]ethyl hexadecanoate (648 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Stas domain-containing protein (10 kDa, Protein from Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1))
Mechanosensitive channel MscS solubilized with DDM in closed conformation
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7onl
Q-score: 0.427
Flegler VJ, Rasmussen A, Borbil K, Boten L, Chen HA, Deinlein H, Halang J, Hellmanzik K, Loffler J, Schmidt V, Makbul C, Kraft C, Hedrich R, Rasmussen T, Bottcher B
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2107095118 Pubmed: 34376558 ]
Cryo-EM structure of E.coli MlaFEB with AMPPNP
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7ch6
Q-score: 0.506
Zhou C, Shi H, Zhang M, Zhou L, Xiao L, Feng S, Im W, Zhou M, Zhang X, Huang Y
J Mol Biol (2021) 433 pp. 166986-166986 [ DOI: doi:10.1016/j.jmb.2021.166986 Pubmed: 33845086 ]
- Phospholipid abc transporter atp-binding protein mlaf (29 kDa, Protein from Escherichia coli (strain K12))
- Lipid asymmetry maintenance protein mlab (10 kDa, Protein from Escherichia coli (strain K12))
- E.coli mlafeb bond with amppnp (Complex from Escherichia coli K12)
- Lipid asymmetry maintenance abc transporter permease subunit mlae (27 kDa, Protein from Escherichia coli (strain K12))
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)