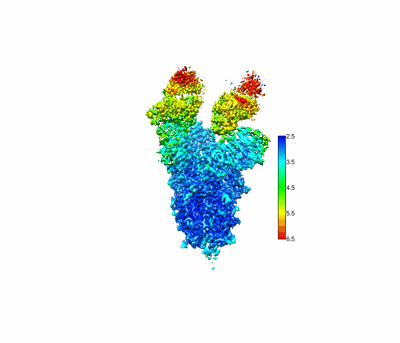
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8khc
Q-score: 0.351
Wang Y, Zhang Z, Yang M, Xiong X, Yan Q, Cao L, Wei P, Zhang Y, Zhang L, Lv K, Chen J, Liu X, Zhao X, Xiao J, Zhang S, Zhu A, Gan M, Zhang J, Cai R, Zhuo J, Zhang Y, Rao H, Qu B, Zhang Y, Chen L, Dai J, Cheng L, Hu Q, Chen Y, Lv H, So RTY, Peiris M, Zhao J, Liu X, Mok CKP, Wang X, Zhao J
Cell Rep (2024) 43 pp. 113653-113653 [ Pubmed: 38175758 DOI: doi:10.1016/j.celrep.2023.113653 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Light chain of 5817 fab (11 kDa, Protein from Homo sapiens)
- Heavy chain of 5817 fab (13 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Spike glycoprotein (124 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron spike in complex with 5817 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
The interface structure of Omicron RBD binding to 5817 Fab
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8khd
Q-score: 0.393
Wang Y, Zhang Z, Yang M, Xiong X, Yan Q, Cao L, Wei P, Zhang Y, Zhang L, Lv K, Chen J, Liu X, Zhao X, Xiao J, Zhang S, Zhu A, Gan M, Zhang J, Cai R, Zhuo J, Zhang Y, Rao H, Qu B, Zhang Y, Chen L, Dai J, Cheng L, Hu Q, Chen Y, Lv H, So RTY, Peiris M, Zhao J, Liu X, Mok CKP, Wang X, Zhao J
Cell Rep (2024) 43 pp. 113653-113653 [ Pubmed: 38175758 DOI: doi:10.1016/j.celrep.2023.113653 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- The interface structure of omicron rbd binding to 5817 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
- Light chain of 5817 (11 kDa, Protein from Homo sapiens)
- Heavy chain of 5817 (13 kDa, Protein from Homo sapiens)
- Spike protein s1 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
BA.2(S375) Spike (S6P)/hACE2 complex
Resolution: 2.92 Å
EM Method: Single-particle
Fitted PDBs: 8wgv
Q-score: 0.376
He X, Zhang X, Wu B, Deng J, Zhang Y, Zhu A, Yuan Y, Lin Y, Chen A, Feng J, Wang X, Wu S, Liu Y, Liu J, Wang Y, Li R, Liang C, Yuan Q, Liang Y, Fang Q, Xi Z, Li W, Liang L, Zhang Z, Tang H, Peng Y, Ke C, Ma X, Cai W, Pan T, Liu B, Deng K, Chen J, Zhao J, Wei X, Chen R, Zhang Y, Zhang H
Nat Immunol (2024) 25 pp. 622-632 [ Pubmed: 38454157 DOI: doi:10.1038/s41590-024-01776-2 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike glycoprotein (138 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Ba.2(s375) spike (s6p)/hace2 complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Angiotensin-converting enzyme 2 (70 kDa, Protein from Homo sapiens)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8wgw
Q-score: 0.541
He X, Zhang X, Wu B, Deng J, Zhang Y, Zhu A, Yuan Y, Lin Y, Chen A, Feng J, Wang X, Wu S, Liu Y, Liu J, Wang Y, Li R, Liang C, Yuan Q, Liang Y, Fang Q, Xi Z, Li W, Liang L, Zhang Z, Tang H, Peng Y, Ke C, Ma X, Cai W, Pan T, Liu B, Deng K, Chen J, Zhao J, Wei X, Chen R, Zhang Y, Zhang H
Nat Immunol (2024) 25 pp. 622-632 [ DOI: doi:10.1038/s41590-024-01776-2 Pubmed: 38454157 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Angiotensin-converting enzyme 2 (70 kDa, Protein from Homo sapiens)
- Rbd complexed with human ace2 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (138 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
Cryo-EM structure of human ZDHHC9/GCP16 complex
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8hf3
Q-score: 0.496
Yang A, Liu S, Zhang Y, Chen J, Fan Y, Wang F, Zou Y, Feng S, Wu J, Hu Q
Nat Struct Mol Biol (2024) 31 pp. 436-446 [ Pubmed: 38182928 DOI: doi:10.1038/s41594-023-01183-5 ]
- Palmitic acid (256 Da, Ligand)
- Golgin subfamily a member 7 (16 kDa, Protein from Homo sapiens)
- Human zdhhc9/gcp16 complex (Complex from Homo sapiens)
- 1,2-dilauroyl-sn-glycero-3-phosphate (535 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Palmitoyltransferase zdhhc9 (41 kDa, Protein from Homo sapiens)
Cryo-EM structure of yeast Erf2/Erf4 complex
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8hfc
Q-score: 0.503
Yang A, Liu S, Zhang Y, Chen J, Fan Y, Wang F, Zou Y, Feng S, Wu J, Hu Q
Nat Struct Mol Biol (2024) 31 pp. 436-446 [ Pubmed: 38182928 DOI: doi:10.1038/s41594-023-01183-5 ]
- Palmitic acid (256 Da, Ligand)
- Palmitoyltransferase erf2 (43 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Ras modification protein erf4 (29 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Erf2/erf4 complex (Complex from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on receptor)
Mao C, Xiao P, Tao XN, Qin J, He QT, Zhang C, Guo SC, Du YQ, Chen LN, Shen DD, Yang ZS, Zhang HQ, Huang SM, He YH, Cheng J, Zhong YN, Shang P, Chen J, Zhang DL, Wang QL, Liu MX, Li GY, Guo Y, Xu HE, Wang C, Zhang C, Feng S, Yu X, Zhang Y, Sun JP
Science (2023) 380 pp. eadd6220-eadd6220 [ Pubmed: 36862765 DOI: doi:10.1126/science.add6220 ]
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on Giq-scFV16 complex)
Mao C, Xiao P, Tao XN, Qin J, He QT, Zhang C, Guo SC, Du YQ, Chen LN, Shen DD, Yang ZS, Zhang HQ, Huang SM, He YH, Cheng J, Zhong YN, Shang P, Chen J, Zhang DL, Wang QL, Liu MX, Li GY, Guo Y, Xu HE, Wang C, Zhang C, Feng S, Yu X, Zhang Y, Sun JP
Science (2023) 380 pp. eadd6220-eadd6220 [ DOI: doi:10.1126/science.add6220 Pubmed: 36862765 ]
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on receptor)
Mao C, Xiao P, Tao XN, Qin J, He QT, Zhang C, Guo SC, Du YQ, Chen LN, Shen DD, Yang ZS, Zhang HQ, Huang SM, He YH, Cheng J, Zhong YN, Shang P, Chen J, Zhang DL, Wang QL, Liu MX, Li GY, Guo Y, Xu HE, Wang C, Zhang C, Feng S, Yu X, Zhang Y, Sun JP
Science (2023) 380 pp. eadd6220-eadd6220 [ Pubmed: 36862765 DOI: doi:10.1126/science.add6220 ]
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on Gil-scFV16 complex)
Mao C, Xiao P, Tao XN, Qin J, He QT, Zhang C, Guo SC, Du YQ, Chen LN, Shen DD, Yang ZS, Zhang HQ, Huang SM, He YH, Cheng J, Zhong YN, Shang P, Chen J, Zhang DL, Wang QL, Liu MX, Li GY, Guo Y, Xu HE, Wang C, Zhang C, Feng S, Yu X, Zhang Y, Sun JP
Science (2023) 380 pp. eadd6220-eadd6220 [ Pubmed: 36862765 DOI: doi:10.1126/science.add6220 ]