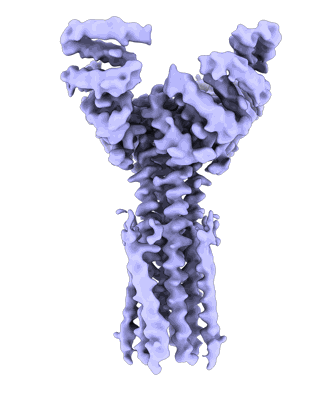
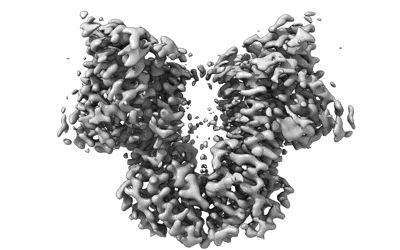
Artificial membrane protein TMHC4_R (ROCKET)
Abramsson M, Corey RA, Skerle J, Persson L, Anden O, Oluwole AO, Howard RJ, Lindahl E, Robinson CV, Strisovsky K, Marklund EG, Drew D, Stansfeld PJ, Landreh M
To Be Published
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 8tlu
Q-score: 0.434
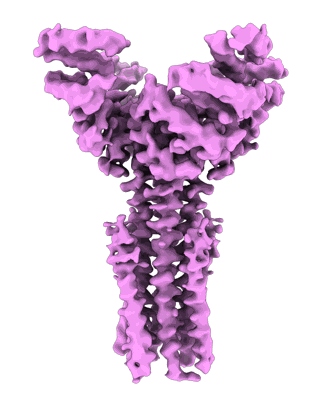
Marmont LS, Orta AK, Baileeves BWA, Sychantha D, Fernandez-Galliano A, Li YE, Greene NG, Corey RA, Stansfeld PJ, Clemons Jr WM, Bernhardt TG
Nat Microbiol (2024) 9 pp. 763-775 [ Pubmed: 38336881 DOI: doi:10.1038/s41564-024-01603-2 ]
- E. coli mray t23p dimer (39 kDa, Cellular component from Escherichia coli K-12)
- Phospho-n-acetylmuramoyl-pentapeptide-transferase (39 kDa, Protein from Escherichia coli K-12)
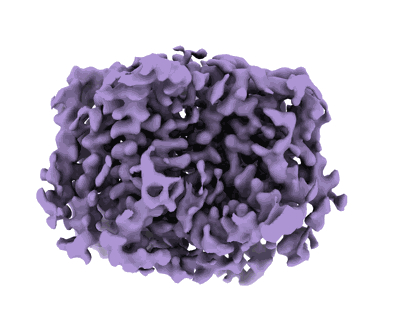
Cryo-EM structure of the fd bacteriophage capsid major coat protein pVIII
Resolution: 3.2 Å
EM Method: Helical reconstruction
Fitted PDBs: 8ch5
Q-score: 0.482
Bohning J, Graham M, Letham SC, Davis LK, Schulze U, Stansfeld PJ, Corey RA, Pearce P, Tarafder AK, Bharat TAM
Nat Commun (2023) 14 pp. 8429-8429 [ DOI: doi:10.1038/s41467-023-44160-8 Pubmed: 38114502 ]
- Major capsid protein pviii (5 kDa, Protein from Escherichia coli)
- Enterobacteria phage fd (Virus from Enterobacteria phage fd)
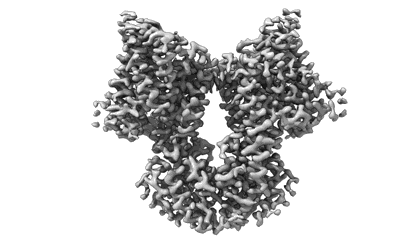
KimA from B. subtilis with nucleotide second-messenger c-di-AMP bound
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8b70
Q-score: 0.536
Fuss MF, Wieferig JP, Corey RA, Hellmich Y, Tascon I, Sousa JS, Stansfeld PJ, Vonck J, Hanelt I
Nat Commun (2023) 14 pp. 3683-3683 [ Pubmed: 37344476 DOI: doi:10.1038/s41467-023-38944-1 ]
- Kima with bound c-di-amp (134 kDa, Complex from Bacillus subtilis)
- Potassium transporter kima (66 kDa, Protein from Bacillus subtilis)
- Potassium ion (39 Da, Ligand)
- (2r,3r,3as,5r,7ar,9r,10r,10as,12r,14ar)-2,9-bis(6-amino-9h-purin-9-yl)octahydro-2h,7h-difuro[3,2-d:3',2'-j][1,3,7,9,2,8 ]tetraoxadiphosphacyclododecine-3,5,10,12-tetrol 5,12-dioxide (658 Da, Ligand)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
Upright KimA dimer with bound c-di-AMP from B. subtilis
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 8b71
Q-score: 0.457
Fuss MF, Wieferig JP, Corey RA, Hellmich Y, Tascon I, Sousa JS, Stansfeld PJ, Vonck J, Hanelt I
Nat Commun (2023) 14 pp. 3683-3683 [ Pubmed: 37344476 DOI: doi:10.1038/s41467-023-38944-1 ]
- (2r,3r,3as,5r,7ar,9r,10r,10as,12r,14ar)-2,9-bis(6-amino-9h-purin-9-yl)octahydro-2h,7h-difuro[3,2-d:3',2'-j][1,3,7,9,2,8 ]tetraoxadiphosphacyclododecine-3,5,10,12-tetrol 5,12-dioxide (658 Da, Ligand)
- Potassium transporter kima (66 kDa, Protein from Bacillus subtilis)
- Upright dimer kima with c-di-amp bound (134 kDa, Complex from Bacillus subtilis)
Single particle cryo-EM of KdpFABC WT (KdpB-Ser162-P) under turnover condition
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hänelt I, Paulino C
Elife (2022) 11 [ DOI: 10.7554/elife.80988 Pubmed: 36255052 ]
Image sets:
- Unaligned movie frames of the KdpFABC WT (KdpB-Ser162-P) under turnover conditions in DDM (Dataset 5)
- Unaligned movie frames of the KdpFABC WT (KdpB-Ser162-P) under turnover conditions in DDM (Dataset 2)
- Unaligned movie frames of the KdpFABC WT (KdpB-Ser162-P) under turnover conditions in DDM (Dataset 3)
- Unaligned movie frames of the KdpFABC WT (KdpB-Ser162-P) under turnover conditions in DDM (Dataset 4)
- Unaligned movie frames of the KdpFABC WT (KdpB-Ser162-P) under turnover conditions in DDM (Dataset 1)
Single particle cryo-EM of KdpFAB(Ser162-P, D307N)C in apo condition
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hänelt I, Paulino C
Elife (2022) 11 [ DOI: 10.7554/elife.80988 Pubmed: 36255052 ]
Image sets:
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 1)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 11)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 2)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 4)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 15)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 6)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 3)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 12)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 14)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 10)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 13)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 8)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 9)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 5)
- Unaligned movie frames of the KdpFAB(Ser162-P, D307N)C without nucleotide in DDM (Dataset 7)
Single particle cryo-EM of KdpFABC WT (KdpB-Ser162-P) in presence of orthovanadate
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hänelt I, Paulino C
Elife (2022) 11 [ DOI: 10.7554/elife.80988 Pubmed: 36255052 ]
Image sets:
Single particle cryo-EM of KdpFAB(S162A)C under turnover condition
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hänelt I, Paulino C
Elife (2022) 11 [ DOI: 10.7554/elife.80988 Pubmed: 36255052 ]
Image sets: