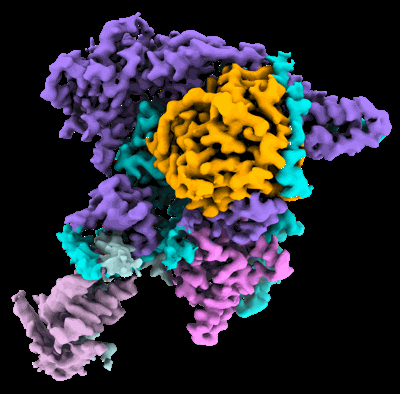
Cryo-EM structure of the histone deacetylase complex Rpd3S
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8hxx
Q-score: 0.453
Nat Struct Mol Biol (2023) 30 pp. 1893-1901 [ Pubmed: 37798513 DOI: doi:10.1038/s41594-023-01121-5 ]
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae)
- Rpd3s histone deacetylase in complex with histone h3 tail (Complex from Saccharomyces cerevisiae)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Rco1 isoform 1 (78 kDa, Protein from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae)
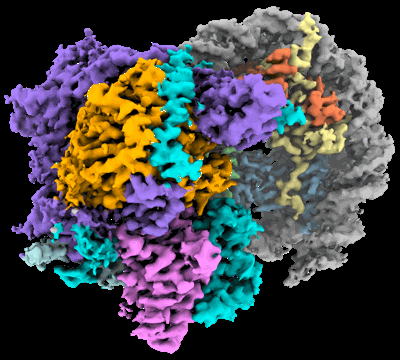
Cryo-EM structure of the histone deacetylase complex Rpd3S in complex with nucleosome
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8hxy
Q-score: 0.44
Nat Struct Mol Biol (2023) 30 pp. 1893-1901 [ Pubmed: 37798513 DOI: doi:10.1038/s41594-023-01121-5 ]
- Dna (352-mer) (108 kDa, DNA from synthetic construct)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae)
- Transcriptional regulatory protein rco1 (78 kDa, Protein from Saccharomyces cerevisiae)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Dna (352-mer) (109 kDa, DNA from synthetic construct)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Rpd3s histone deacetylase in complex with nucleosome (680 kDa, Complex from Saccharomyces cerevisiae)
Cryo-EM structure of Eaf3 CHD in complex with nucleosome
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8hxz
Q-score: 0.326
Nat Struct Mol Biol (2023) 30 pp. 1893-1901 [ Pubmed: 37798513 DOI: doi:10.1038/s41594-023-01121-5 ]
- Dna (352-mer) (108 kDa, DNA from synthetic construct)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Dna (352-mer) (109 kDa, DNA from synthetic construct)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Eaf3 chd in complex with nucleosome (150 kDa, Complex from Saccharomyces)
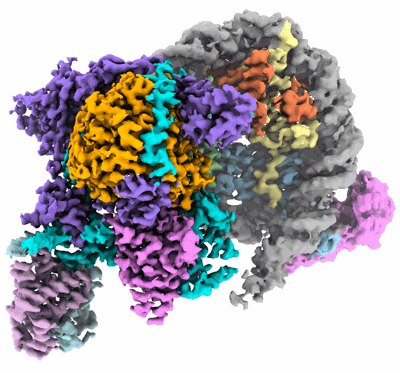
Composite cryo-EM structure of the histone deacetylase complex Rpd3S in complex with nucleosome
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8hy0
Q-score: 0.463
Nat Struct Mol Biol (2023) 30 pp. 1893-1901 [ Pubmed: 37798513 DOI: doi:10.1038/s41594-023-01121-5 ]
- Rpd3s histone deacetylase in complex with nucleosome (800 kDa, Complex from Saccharomyces cerevisiae)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae)
- Rco1 isoform 1 (78 kDa, Protein from Saccharomyces cerevisiae)
- Dna (352-mer) (108 kDa, DNA from synthetic construct)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae)
- Dna (352-mer) (109 kDa, DNA from synthetic construct)
Resolution: 3.66 Å
EM Method: Single-particle
Fitted PDBs: 8hih
Q-score: 0.202
Shi J, Feng Z, Xu J, Li F, Zhang Y, Wen A, Wang F, Song Q, Wang L, Cui H, Tong S, Chen P, Zhu Y, Zhao G, Wang S, Feng Y, Lin W
PNAS (2023) 120 pp. e2300282120-e2300282120 [ Pubmed: 37216560 DOI: doi:10.1073/pnas.2300282120 ]
- Dna-directed rna polymerase subunit beta (130 kDa, Protein from Mycobacterium tuberculosis)
- Dna-directed rna polymerase subunit omega (11 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Transcriptional regulatory protein (27 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Dna-directed rna polymerase subunit beta' (146 kDa, Protein from Mycobacterium tuberculosis)
- Zinc ion (65 Da, Ligand)
- Mycobacterium tuberculosis transcription initiation complex with transcription factor glnr (Complex from Mycobacterium tuberculosis)
- Rna polymerase sigma factor siga (57 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Dna-directed rna polymerase subunit alpha (37 kDa, Protein from Mycobacterium tuberculosis)
- Magnesium ion (24 Da, Ligand)
- Non-template strand dna of amtb promoter (33 kDa, DNA from Mycobacterium tuberculosis)
- Template strand dna of amtb promoter (33 kDa, DNA from Mycobacterium tuberculosis)
CryoEM structure of the C. sordellii lethal toxin TcsL in complex with SEMA6A
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 6wts
Q-score: 0.463
Lee H, Beilhartz GL, Kucharska I, Raman S, Cui H, Lam MHY, Liang H, Rubinstein JL, Schramek D, Julien JP, Melnyk RA, Taipale M
Cell (2020) 182 pp. 345- [ Pubmed: 32589945 DOI: doi:10.1016/j.cell.2020.06.005 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Tcsl (Complex from Paeniclostridium sordellii)
- Sema6a (63 kDa, Protein from Homo sapiens)
- Tcsl (63 kDa, Protein from Paeniclostridium sordellii)
- Sema6a (Complex from Homo sapiens)
- Complex of sema6a with tcsl (Complex)