Cryo-EM structure of CII-dependent transcription activation complex
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8igr
Q-score: 0.52
Zhao M, Gao B, Wen A, Feng Y, Lu YQ
Structure (2023) 31 pp. 968- [ DOI: doi:10.1016/j.str.2023.05.008 Pubmed: 37269829 ]
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli (strain K12))
- Template strand dna (26 kDa, DNA from Escherichia coli (strain K12))
- Zinc ion (65 Da, Ligand)
- Rna polymerase sigma factor rpod (70 kDa, Protein from Escherichia coli (strain K12))
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli (strain K12))
- Transcriptional activator ii (11 kDa, Protein from Escherichia phage Lambda)
- Dna-directed rna polymerase subunit omega (10 kDa, Protein from Escherichia coli (strain K12))
- Nontemplate strand dna (26 kDa, DNA from Escherichia coli (strain K12))
- Magnesium ion (24 Da, Ligand)
- Dna-directed rna polymerase subunit beta' (155 kDa, Protein from Escherichia coli (strain K12))
- Cii-dependent transcription activation complex (Complex from Escherichia coli)
Cryo-EM structure of RNAP-promoter open complex at lambda promoter PRE
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8igs
Q-score: 0.469
Zhao M, Gao B, Wen A, Feng Y, Lu YQ
Structure (2023) 31 pp. 968- [ DOI: doi:10.1016/j.str.2023.05.008 Pubmed: 37269829 ]
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli (strain K12))
- Template strand dna (26 kDa, DNA from Escherichia coli (strain K12))
- Non-template strand dna (26 kDa, DNA from Escherichia coli (strain K12))
- Zinc ion (65 Da, Ligand)
- Rna polymerase sigma factor rpod (70 kDa, Protein from Escherichia coli (strain K12))
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli (strain K12))
- Rnap-promoter open complex (Complex from Escherichia coli (strain K12))
- Dna-directed rna polymerase subunit omega (10 kDa, Protein from Escherichia coli (strain K12))
- Magnesium ion (24 Da, Ligand)
- Dna-directed rna polymerase subunit beta' (155 kDa, Protein from Escherichia coli (strain K12))
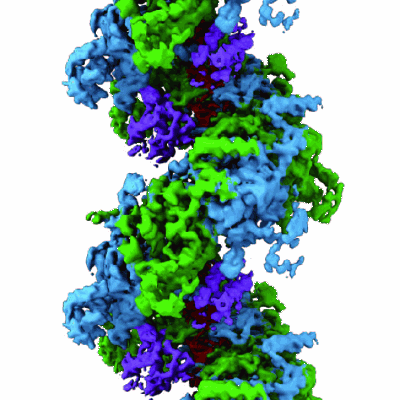
Structure of DinI in complex with RecA filament
Resolution: 3.26 Å
EM Method: Helical reconstruction
Fitted PDBs: 7ywa
Q-score: 0.455
Gao B, Liang L, Su L, Wen A, Zhou C, Feng Y
PNAS (2023) 120 pp. e2217493120-e2217493120 [ Pubmed: 36598938 DOI: doi:10.1073/pnas.2217493120 ]
- Dna damage-inducible protein i (8 kDa, Protein from Escherichia coli)
- Reca-dini complex (Complex from Escherichia coli)
- Protein reca (38 kDa, Protein from Escherichia coli)
- Reca-dini (Complex)
- Dna (5'-d(p*tp*tp*tp*tp*tp*t)-3') (1 kDa, DNA from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Dna (Complex)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
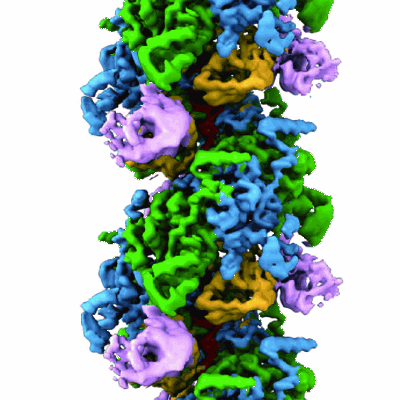
Structure of LexA in complex with RecA filament
Resolution: 3.31 Å
EM Method: Helical reconstruction
Fitted PDBs: 8gms
Q-score: 0.478
Gao B, Liang L, Su L, Wen A, Zhou C, Feng Y
PNAS (2023) 120 pp. e2217493120-e2217493120 [ Pubmed: 36598938 DOI: doi:10.1073/pnas.2217493120 ]
- Protein reca (38 kDa, Protein from Escherichia coli)
- Reca-lexa complex (Complex from Escherichia coli)
- Dna (5'-d(p*tp*tp*tp*tp*tp*t)-3') (1 kDa, DNA from Escherichia coli)
- Lexa repressor (14 kDa, Protein from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Dna (Complex)
- Reca-lexa (Complex)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
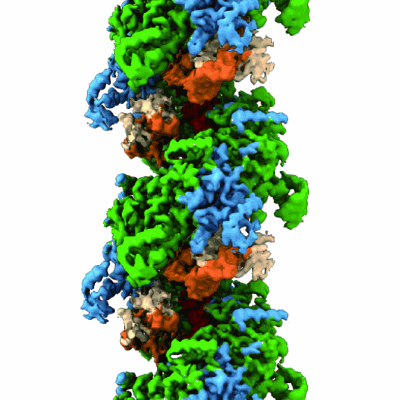
Structure of UmuD in complex with RecA filament
Resolution: 3.31 Å
EM Method: Helical reconstruction
Fitted PDBs: 8gmt
Q-score: 0.493
Gao B, Liang L, Su L, Wen A, Zhou C, Feng Y
PNAS (2023) 120 pp. e2217493120-e2217493120 [ Pubmed: 36598938 DOI: doi:10.1073/pnas.2217493120 ]
- Protein reca (38 kDa, Protein from Escherichia coli)
- Dna polymerase v subunit umud (15 kDa, Protein from Escherichia coli)
- Reca-umud complex (Complex from Escherichia coli)
- Dna (5'-d(p*tp*tp*tp*tp*tp*t)-3') (1 kDa, DNA from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
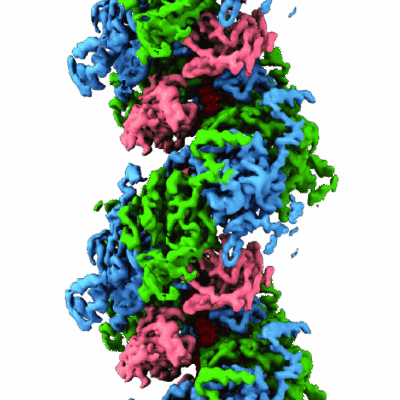
Structure of lambda repressor in complex with RecA filament
Resolution: 2.78 Å
EM Method: Helical reconstruction
Fitted PDBs: 8gmu
Q-score: 0.585
Gao B, Liang L, Su L, Wen A, Zhou C, Feng Y
PNAS (2023) 120 pp. e2217493120-e2217493120 [ Pubmed: 36598938 DOI: doi:10.1073/pnas.2217493120 ]
- Repressor protein ci (14 kDa, Protein from Escherichia phage Lambda)
- Protein reca (38 kDa, Protein from Escherichia coli)
- Dna (5'-d(p*tp*tp*tp*tp*tp*t)-3') (1 kDa, DNA from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Reca-ci complex (Complex from Escherichia phage Lambda)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
Cryo-EM structure of Arabidopsis DCL1 in complex with pri-miRNA 166f
Resolution: 4.6 Å
EM Method: Single-particle
Fitted PDBs: 7eld
Q-score: 0.157
Wei X, Ke H, Wen A, Gao B, Shi J, Feng Y
Nat Plants (2021) 7 pp. 1389-1396 [ Pubmed: 34593993 DOI: doi:10.1038/s41477-021-01000-1 ]
- Binary complex of dcl1 in complex with pri-mirna 166f (Complex from Arabidopsis thaliana)
- Pri-mirna 166f (47 kDa, RNA from Arabidopsis thaliana)
- Endoribonuclease dicer homolog 1 (213 kDa, Protein from Arabidopsis thaliana)
Cryo-EM structure of Arabidopsis DCL1 in complex with pre-miRNA 166f
Resolution: 4.9 Å
EM Method: Single-particle
Fitted PDBs: 7ele
Q-score: 0.186
Wei X, Ke H, Wen A, Gao B, Shi J, Feng Y
Nat Plants (2021) 7 pp. 1389-1396 [ Pubmed: 34593993 DOI: doi:10.1038/s41477-021-01000-1 ]
- Binary complex of dcl1 in complex with pre-mirna 166f (Complex from Arabidopsis thaliana)
- Pre-mirna 166f (28 kDa, RNA from Arabidopsis thaliana)
- Endoribonuclease dicer homolog 1 (213 kDa, Protein from Arabidopsis thaliana)
CryoEM structure of gp55-dependent RNA polymerase-promoter open complex
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7d7c
Q-score: 0.444
Shi J, Wen A, Jin S, Gao B, Huang Y, Feng Y
Nat Commun (2021) 12 pp. 1131-1131 [ DOI: doi:10.1038/s41467-021-21392-0 Pubmed: 33602900 ]
- Dna (nontemplate strand) (18 kDa, DNA from Escherichia coli)
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli)
- Gp55-dependent rna polymerase-promoter open complex (Complex from Thermus thermophilus)
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli 1-392-07 S4 C3)
- Dna (template strand) (18 kDa, DNA from Escherichia coli)
- Dna-directed rna polymerase subunit beta' (155 kDa, Protein from Escherichia coli)
- Gp55 (21 kDa, Protein from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
CryoEM structure of gp45-dependent transcription activation complex
Resolution: 4.5 Å
EM Method: Single-particle
Fitted PDBs: 7d7d
Q-score: 0.316
Shi J, Wen A, Jin S, Gao B, Huang Y, Feng Y
Nat Commun (2021) 12 pp. 1131-1131 [ DOI: doi:10.1038/s41467-021-21392-0 Pubmed: 33602900 ]
- Dna (template strand) (18 kDa, DNA from Escherichia virus T4)
- Dna polymerase clamp (25 kDa, Protein from Enterobacteria phage T4)
- Magnesium ion (24 Da, Ligand)
- Gp45-dependent transcription activation complex (Complex from Thermus thermophilus)
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli 1-392-07 S4 C3)
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli)
- Rna polymerase-associated protein gp33 (15 kDa, Protein from Enterobacteria phage T4)
- Dna-directed rna polymerase subunit omega (10 kDa, Protein from Escherichia coli)
- Dna-directed rna polymerase subunit beta' (155 kDa, Protein from Escherichia coli)
- Dna (nontemplate strand) (18 kDa, DNA from Escherichia virus T4)
- Zinc ion (65 Da, Ligand)
- Gp55 (21 kDa, Protein from Escherichia virus T4)