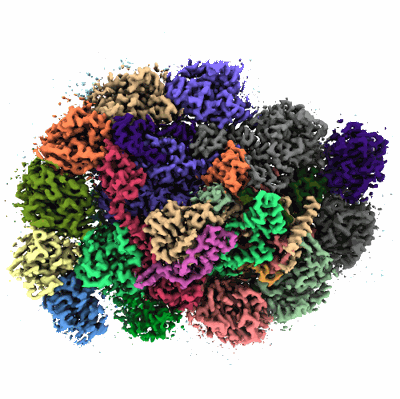
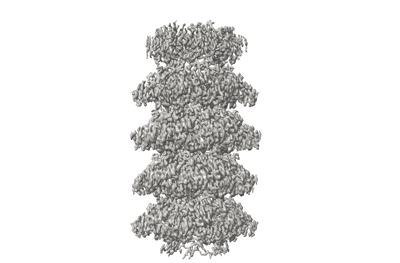
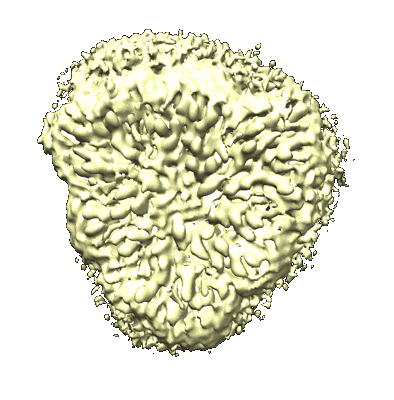
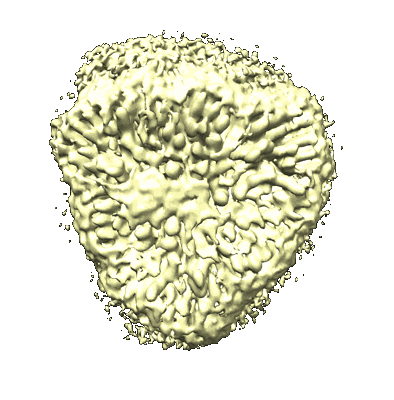
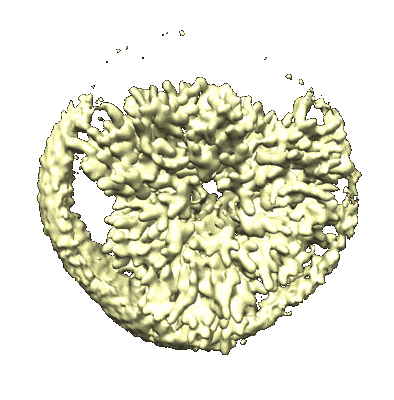
Cryo-EM structure of Symbiodinium photosystem I
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8jjr
Q-score: 0.575
Zhao LS, Wang N, Li K, Li CY, Guo JP, He FY, Liu GM, Chen XL, Gao J, Liu LN, Zhang YZ
Nat Commun (2024) 15 pp. 2392-2392 [ DOI: doi:10.1038/s41467-024-46791-x Pubmed: 38493166 ]
- Pcpi-1 (28 kDa, Protein from Symbiodinium sp.)
- Psat (13 kDa, Protein from Symbiodinium sp.)
- Pcp-11 (21 kDa, Protein from Symbiodinium sp.)
- Psai (19 kDa, Protein from Symbiodinium sp.)
- Phylloquinone (450 Da, Ligand)
- Psad (32 kDa, Protein from Symbiodinium sp.)
- Pcpi-5 (26 kDa, Protein from Symbiodinium sp.)
- Chlorophyll a (893 Da, Ligand)
- Pcpi-3 (27 kDa, Protein from Symbiodinium sp.)
- [(1~{s},5~{r})-3,3,5-trimethyl-5-oxidanyl-4-[(3~{e},5~{e},7~{e},9~{e},11~{e},13~{e},15~{e},17~{e})-3,7,12,16-tetramethyl-18-[(1~{s},4~{s},6~{r})-2,2,6-trimethyl-4-oxidanyl-7-oxabicyclo[4.1.0]heptan-1-yl]octadeca-1,3,5,7,9,11,13,15,17-nonaenylidene]cyclohexyl] ethanoate (642 Da, Ligand)
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- Photosystem i of symbiodinium (Complex from Symbiodinium sp.)
- Pcpi-7 (27 kDa, Protein from Symbiodinium sp.)
- Chlorophyll c2 (608 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Pcpi-2 (15 kDa, Protein from Symbiodinium sp.)
- Beta-carotene (536 Da, Ligand)
- Psam (14 kDa, Protein from Symbiodinium sp.)
- Peridinin (630 Da, Ligand)
- Pcpi-10 (27 kDa, Protein from Symbiodinium sp.)
- Psal (38 kDa, Protein from Symbiodinium sp.)
- Pcpi-13 (23 kDa, Protein from Symbiodinium sp.)
- Pcpi-6 (19 kDa, Protein from Symbiodinium sp.)
- Psaf (29 kDa, Protein from Symbiodinium sp.)
- Psau (24 kDa, Protein from Symbiodinium sp.)
- (3s,3'r,5r,6s,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,beta-carotene-3,3'-diol (582 Da, Ligand)
- Pcpi-9 (22 kDa, Protein from Symbiodinium sp.)
- Psab (74 kDa, Protein from Symbiodinium sp.)
- Psaa (76 kDa, Protein from Symbiodinium sp.)
- Pcpi-8 (30 kDa, Protein from Symbiodinium sp.)
- 1,2-di-o-acyl-3-o-[6-deoxy-6-sulfo-alpha-d-glucopyranosyl]-sn-glycerol (795 Da, Ligand)
- Pcpi-12 (21 kDa, Protein from Symbiodinium sp.)
- Dodecyl-alpha-d-maltoside (510 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Digalactosyl diacyl glycerol (dgdg) (949 Da, Ligand)
- Psac (16 kDa, Protein from Symbiodinium sp.)
- Psae (13 kDa, Protein from Symbiodinium sp.)
- Psaj (15 kDa, Protein from Symbiodinium sp.)
- Psar (17 kDa, Protein from Symbiodinium sp.)
- Pcpi-4 (24 kDa, Protein from Symbiodinium sp.)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8k98
Q-score: 0.486
Yin H, Li X, Wang X, Zhang C, Gao J, Yu G, He Q, Yang J, Liu X, Wei Y, Li Z, Zhang H
Nat Commun (2024) 15 pp. 2692-2692 [ Pubmed: 38538592 DOI: doi:10.1038/s41467-024-47030-z ]
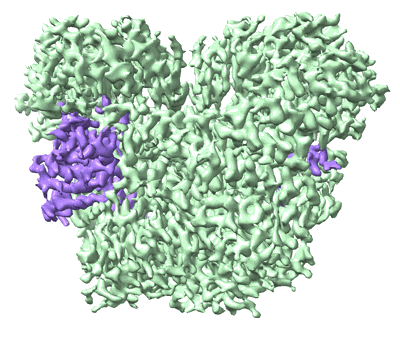
- Cryo-em structure of a protein (Complex from Bacillus halotolerans)
- A protein (29 kDa, Protein from Bacillus halotolerans)
- A protein (118 kDa, Protein from Bacillus halotolerans)
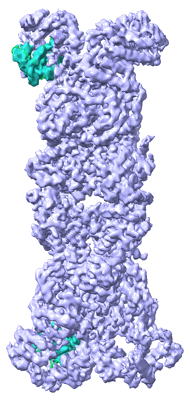
Cryo-EM structure of tail tube protein
Resolution: 3.11 Å
EM Method: Single-particle
Fitted PDBs: 8xkn
Q-score: 0.542
Yin H, Li X, Wang X, Zhang C, Gao J, Yu G, He Q, Yang J, Liu X, Wei Y, Li Z, Zhang H
Nat Commun (2024) 15 pp. 2692-2692 [ Pubmed: 38538592 DOI: doi:10.1038/s41467-024-47030-z ]
- A protein (29 kDa, Protein from Bacillus halotolerans)
- Cryo-em structure of tail tube protein (Complex from Bacillus halotolerans)
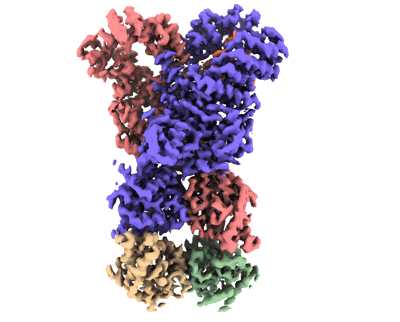
Cryo-EM structure of unprotonated LHCII in detergent solution at high pH value
Resolution: 2.59 Å
EM Method: Single-particle
Fitted PDBs: 8iwx
Q-score: 0.466
Ruan M, Li H, Zhang Y, Zhao R, Zhang J, Wang Y, Gao J, Wang Z, Wang Y, Sun D, Ding W, Weng Y
Nat Plants (2023) 9 pp. 1547-1557 [ Pubmed: 37653340 DOI: doi:10.1038/s41477-023-01500-2 ]
- Chlorophyll b (907 Da, Ligand)
- (3r,3'r,6s)-4,5-didehydro-5,6-dihydro-beta,beta-carotene-3,3'-diol (568 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Chlorophyll a-b binding protein, chloroplastic (23 kDa, Protein from Spinacia oleracea)
- Lhcii in detergent solution at high ph value (120 kDa, Complex from Spinacia oleracea)
- (3s,5r,6s,3's,5'r,6's)-5,6,5',6'-diepoxy-5,6,5',6'- tetrahydro-beta,beta-carotene-3,3'-diol (600 Da, Ligand)
- (1r,3r)-6-{(3e,5e,7e,9e,11e,13e,15e,17e)-18-[(1s,4r,6r)-4-hydroxy-2,2,6-trimethyl-7-oxabicyclo[4.1.0]hept-1-yl]-3,7,12,16-tetramethyloctadeca-1,3,5,7,9,11,13,15,17-nonaenylidene}-1,5,5-trimethylcyclohexane-1,3-diol (600 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)
Cryo-EM structure of protonated LHCII in detergent solution at low pH value
Resolution: 2.68 Å
EM Method: Single-particle
Fitted PDBs: 8iwy
Q-score: 0.572
Ruan M, Li H, Zhang Y, Zhao R, Zhang J, Wang Y, Gao J, Wang Z, Wang Y, Sun D, Ding W, Weng Y
Nat Plants (2023) 9 pp. 1547-1557 [ Pubmed: 37653340 DOI: doi:10.1038/s41477-023-01500-2 ]
- Chlorophyll b (907 Da, Ligand)
- Lhcii in detergent solution at low ph value (120 kDa, Complex from Spinacia oleracea)
- (3r,3'r,6s)-4,5-didehydro-5,6-dihydro-beta,beta-carotene-3,3'-diol (568 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Chlorophyll a-b binding protein, chloroplastic (23 kDa, Protein from Spinacia oleracea)
- (3s,5r,6s,3's,5'r,6's)-5,6,5',6'-diepoxy-5,6,5',6'- tetrahydro-beta,beta-carotene-3,3'-diol (600 Da, Ligand)
- (1r,3r)-6-{(3e,5e,7e,9e,11e,13e,15e,17e)-18-[(1s,4r,6r)-4-hydroxy-2,2,6-trimethyl-7-oxabicyclo[4.1.0]hept-1-yl]-3,7,12,16-tetramethyloctadeca-1,3,5,7,9,11,13,15,17-nonaenylidene}-1,5,5-trimethylcyclohexane-1,3-diol (600 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)
Cryo-EM structure of unprotonated LHCII in detergent solution at low pH value
Resolution: 2.52 Å
EM Method: Single-particle
Fitted PDBs: 8iwz
Q-score: 0.563
Ruan M, Li H, Zhang Y, Zhao R, Zhang J, Wang Y, Gao J, Wang Z, Wang Y, Sun D, Ding W, Weng Y
Nat Plants (2023) 9 pp. 1547-1557 [ Pubmed: 37653340 DOI: doi:10.1038/s41477-023-01500-2 ]
- Lhcii in detergent solution at low ph value (120 kDa, Complex from Spinacia oleracea)
- (1r,3r)-6-{(3e,5e,7e,9e,11e,13e,15e,17e)-18-[(1s,4r,6r)-4-hydroxy-2,2,6-trimethyl-7-oxabicyclo[4.1.0]hept-1-yl]-3,7,12,16-tetramethyloctadeca-1,3,5,7,9,11,13,15,17-nonaenylidene}-1,5,5-trimethylcyclohexane-1,3-diol (600 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)
- Chlorophyll a-b binding protein, chloroplastic (23 kDa, Protein from Spinacia oleracea)
- Chlorophyll b (907 Da, Ligand)
- (3s,5r,6s,3's,5'r,6's)-5,6,5',6'-diepoxy-5,6,5',6'- tetrahydro-beta,beta-carotene-3,3'-diol (600 Da, Ligand)
- (3r,3'r,6s)-4,5-didehydro-5,6-dihydro-beta,beta-carotene-3,3'-diol (568 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
Cryo-EM structure of unprotonated LHCII nanodisc at high pH value
Resolution: 2.64 Å
EM Method: Single-particle
Fitted PDBs: 8ix0
Q-score: 0.541
Ruan M, Li H, Zhang Y, Zhao R, Zhang J, Wang Y, Gao J, Wang Z, Wang Y, Sun D, Ding W, Weng Y
Nat Plants (2023) 9 pp. 1547-1557 [ Pubmed: 37653340 DOI: doi:10.1038/s41477-023-01500-2 ]
- Chlorophyll b (907 Da, Ligand)
- (3r,3'r,6s)-4,5-didehydro-5,6-dihydro-beta,beta-carotene-3,3'-diol (568 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Chlorophyll a-b binding protein, chloroplastic (23 kDa, Protein from Spinacia oleracea)
- Lhcii nanodsic at high ph value (120 kDa, Complex from Spinacia oleracea)
- (3s,5r,6s,3's,5'r,6's)-5,6,5',6'-diepoxy-5,6,5',6'- tetrahydro-beta,beta-carotene-3,3'-diol (600 Da, Ligand)
- (1r,3r)-6-{(3e,5e,7e,9e,11e,13e,15e,17e)-18-[(1s,4r,6r)-4-hydroxy-2,2,6-trimethyl-7-oxabicyclo[4.1.0]hept-1-yl]-3,7,12,16-tetramethyloctadeca-1,3,5,7,9,11,13,15,17-nonaenylidene}-1,5,5-trimethylcyclohexane-1,3-diol (600 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)