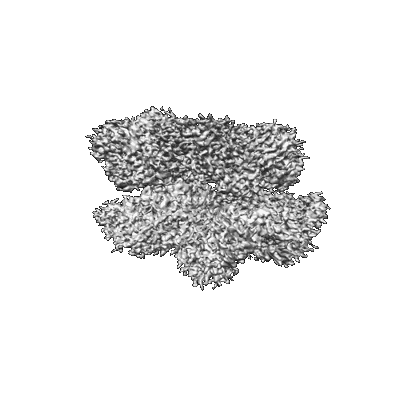
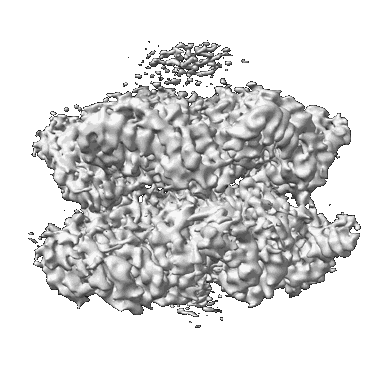
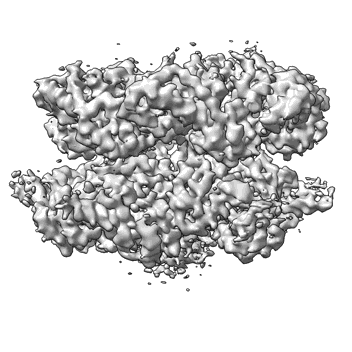
CryoEM structure of the N-Terminal deleted Rix7 AAA-ATPase
Resolution: 2.88 Å
EM Method: Single-particle
Fitted PDBs: 7swl
Q-score: 0.546
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ DOI: doi:10.1093/pnasnexus/pgac118 Pubmed: 36090660 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Cryoem structure of the n-terminal-deleted rix7 aaa-atpase (100 kDa, Complex from Chaetomium thermophilum)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Rix7 (69 kDa, Protein from Chaetomium thermophilum)
- Polyleucine (2 kDa, Protein from synthetic construct)
CryoEM structure of the crosslinked Rix7 AAA-ATPase
Resolution: 3.67 Å
EM Method: Single-particle
Fitted PDBs: 7t0v
Q-score: 0.381
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ DOI: doi:10.1093/pnasnexus/pgac118 Pubmed: 36090660 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cryoem structure of the crosslinked rix7 aaa-atpase (100 kDa, Complex from Chaetomium thermophilum)
- Magnesium ion (24 Da, Ligand)
- Polyvaline (2 kDa, Protein from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Rix7 (89 kDa, Protein from Chaetomium thermophilum)
CryoEM structure of the Rix7 D2 Walker B mutant
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 7t3i
Q-score: 0.38
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ Pubmed: 36090660 DOI: doi:10.1093/pnasnexus/pgac118 ]
- Substrate peptide (2 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cryoem structure of rix7 d2 walker b mutant (Complex from Chaetomium thermophilum)
- Phosphate ion (94 Da, Ligand)
- Rix7 (89 kDa, Protein from Chaetomium thermophilum)
Nucleotide bound SARS-CoV-2 Nsp15
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7k0r
Q-score: 0.431
Pillon MC, Frazier MN, Dillard LB, Williams JG, Kocaman S, Krahn JM, Perera L, Hayne CK, Gordon J, Stewart ZD, Sobhany M, Deterding LJ, Hsu AL, Dandey VP, Borgnia MJ, Stanley RE
Nat Commun (2021) 12 pp. 636-636 [ DOI: doi:10.1038/s41467-020-20608-z Pubmed: 33504779 ]
- Nucleotide bound complex of sars-cov-2 nsp15 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Uridine-5'-monophosphate (324 Da, Ligand)
- Uridylate-specific endoribonuclease (40 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphate ion (94 Da, Ligand)
Pillon MC, Frazier MN, Dillard LB, Williams JG, Kocaman S, Krahn JM, Perera L, Hayne CK, Gordon J, Stewart ZD, Sobhany M, Deterding LJ, Hsu AL, Dandey VP, Borgnia MJ, Stanley RE
Nat Commun (2021) 12 pp. 636-636 [ Pubmed: 33504779 DOI: doi:10.1038/s41467-020-20608-z ]
SARS-CoV-2 Nsp15 H235A variant APO-state, dataset ii
Pillon MC, Frazier MN, Dillard LB, Williams JG, Kocaman S, Krahn JM, Perera L, Hayne CK, Gordon J, Stewart ZD, Sobhany M, Deterding LJ, Hsu AL, Dandey VP, Borgnia MJ, Stanley RE
Nat Commun (2021) 12 pp. 636-636 [ Pubmed: 33504779 DOI: doi:10.1038/s41467-020-20608-z ]
SARS-CoV-2 Nsp15 H235A APO-state, dataset i
Pillon MC, Frazier MN, Dillard LB, Williams JG, Kocaman S, Krahn JM, Perera L, Hayne CK, Gordon J, Stewart ZD, Sobhany M, Deterding LJ, Hsu AL, Dandey VP, Borgnia MJ, Stanley RE
Nat Commun (2021) 12 pp. 636-636 [ DOI: doi:10.1038/s41467-020-20608-z Pubmed: 33504779 ]