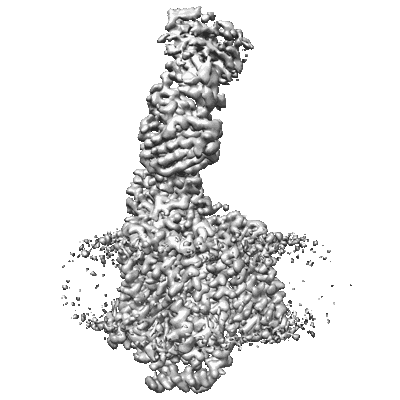
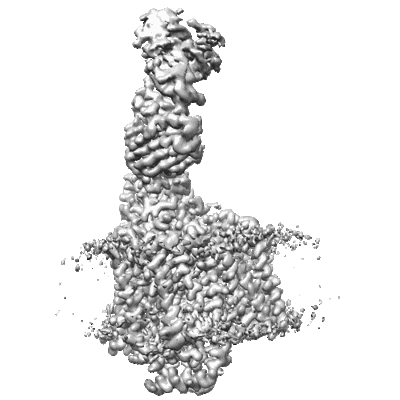
Cryo-EM structure of Cytochrome bo3 from Escherichia coli, apo structure with DMSO
Resolution: 3.09 Å
EM Method: Single-particle
Fitted PDBs: 7xmc
Q-score: 0.526
Nishida Y, Yanagisawa S, Morita R, Shigematsu H, Shinzawa-Itoh K, Yuki H, Ogasawara S, Shimuta K, Iwamoto T, Nakabayashi C, Matsumura W, Kato H, Gopalasingam C, Nagao T, Qaqorh T, Takahashi Y, Yamazaki S, Kamiya K, Harada R, Mizuno N, Takahashi H, Akeda Y, Ohnishi M, Ishii Y, Kumasaka T, Murata T, Muramoto K, Tosha T, Shiro Y, Honma T, Shigeta Y, Kubo M, Takashima S, Shintani Y
Nat Commun (2022) 13 pp. 7591-7591 [ Pubmed: 36481732 DOI: doi:10.1038/s41467-022-34771-y ]
- Heme o (838 Da, Ligand)
- Copper (ii) ion (63 Da, Ligand)
- Ubiquinol oxidase subunit 2 (36 kDa, Protein from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 1 (74 kDa, Protein from Escherichia coli)
- Cytochrome bo3 ubiquinol oxidase (144 kDa, Complex from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 4 (12 kDa, Protein from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 3 (22 kDa, Protein from Escherichia coli)
- Unknown atom or ion (744 Da, Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Protoporphyrin ix containing fe (616 Da, Ligand)
Resolution: 2.99 Å
EM Method: Single-particle
Fitted PDBs: 7xmd
Q-score: 0.535
Nishida Y, Yanagisawa S, Morita R, Shigematsu H, Shinzawa-Itoh K, Yuki H, Ogasawara S, Shimuta K, Iwamoto T, Nakabayashi C, Matsumura W, Kato H, Gopalasingam C, Nagao T, Qaqorh T, Takahashi Y, Yamazaki S, Kamiya K, Harada R, Mizuno N, Takahashi H, Akeda Y, Ohnishi M, Ishii Y, Kumasaka T, Murata T, Muramoto K, Tosha T, Shiro Y, Honma T, Shigeta Y, Kubo M, Takashima S, Shintani Y
Nat Commun (2022) 13 pp. 7591-7591 [ Pubmed: 36481732 DOI: doi:10.1038/s41467-022-34771-y ]
- Heme o (838 Da, Ligand)
- Copper (ii) ion (63 Da, Ligand)
- Ubiquinol oxidase subunit 2 (36 kDa, Protein from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 1 (74 kDa, Protein from Escherichia coli)
- Cytochrome bo3 ubiquinol oxidase (144 kDa, Complex from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 4 (12 kDa, Protein from Escherichia coli)
- Cytochrome bo(3) ubiquinol oxidase subunit 3 (22 kDa, Protein from Escherichia coli)
- Unknown atom or ion (Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Methyl 3-oxidanyl-5-[oxidanyl(oxidanylidene)-$l^{4}-azanyl]-1-benzothiophene-2-carboxylate (253 Da, Ligand)
- Protoporphyrin ix containing fe (616 Da, Ligand)
Cryo-EM reconstruction of equine apoferritin (unmodified)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of equine apoferritin (1 mM PEG4)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of equine apoferritin (4 mM PEG4)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of equine apoferritin (1 mM PEG8)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of equine apoferritin (4 mM PEG8)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of E.coli beta-galactosidase (unmodified)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of E.coli beta-galactosidase (2 mM PEG4)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]
Cryo-EM reconstruction of E.coli beta-galactosidase (2 mM PEG8)
Zhang Z, Shigematsu H, Shimizu T, Ohto U
Structure (2021) 29 pp. 1192-1199.e4 [ Pubmed: 34048698 DOI: doi:10.1016/j.str.2021.05.004 ]