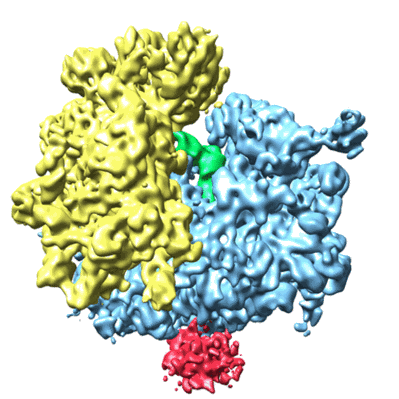
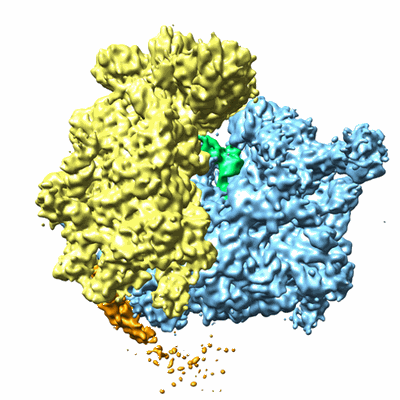
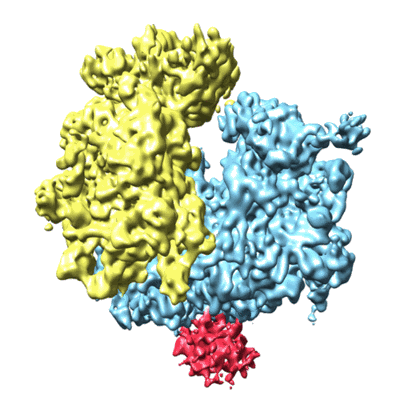
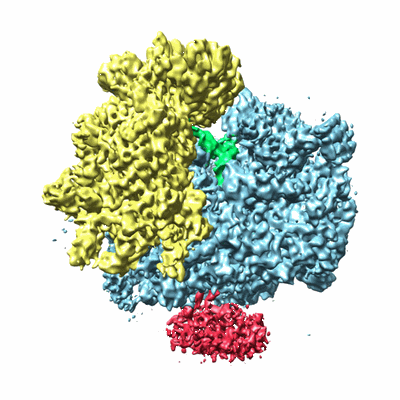
Cryo-electron microscopy structure of the Trypanosoma brucei 80S ribosome
Resolution: 5.57 Å
EM Method: Single-particle
Fitted PDBs: 4v8m
Q-score: 0.239
Hashem Y, des Georges A, Fu J, Buss SN, Jossinet F, Jobe A, Zhang Q, Liao HY, Grassucci RA, Bajaj C, Westhof E, Madison-Antenucci S, Frank J
Nature (2013) 494 pp. 385-389 [ Pubmed: 23395961 DOI: doi:10.1038/nature11872 ]
- 80s ribosome (3 MDa, Complex from Trypanosoma brucei)
- Trypanosoma brucei 80s ribosome (3 MDa, Sample)
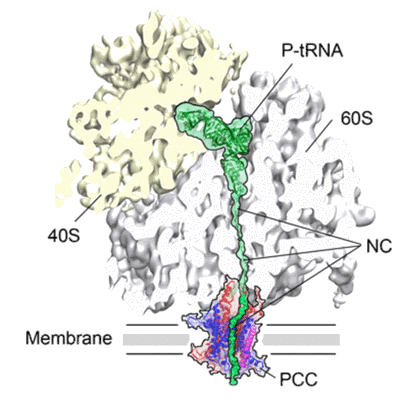
Cryo-EM structure of the programmed yeast 80 ribosome bound the Ssh1 complex
Becker T, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Mandon EC, Beckmann R
Science (2009) 326 pp. 1369-1373 [ Pubmed: 19933108 DOI: doi:10.1126/science.1178535 ]
- Ssh1 complex (71 kDa, Ligand from Saccharomyces cerevisiae)
- Yeast 80s ribosome bound to the yeast ssh1 complex (4 MDa, Complex)
- This map represents a yeast 80s ribosome in the unratcheted conformation with the yeast ssh1 complex bound after sorting the entire dataset against an empty (ratcheted) yeast 80s ribosome (4 MDa, Sample)
Resolution: 8.6 Å
EM Method: Single-particle
Fitted PDBs: 2ww9
Q-score: 0.078
Becker T, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Mandon EC, Beckmann R
Science (2009) 326 pp. 1369-1373 [ Pubmed: 19933108 DOI: doi:10.1126/science.1178535 ]
Becker T, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Mandon EC, Beckmann R
Science (2009) 326 pp. 1369-1373 [ DOI: doi:10.1126/science.1178535 Pubmed: 19933108 ]
Cryo-EM structures of the idle yeast Ssh1 complex bound to the yeast 80S ribosome
Resolution: 8.8 Å
EM Method: Single-particle
Fitted PDBs: 4v6i, 3izd
Q-score: 0.0565
Becker T, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Mandon EC, Beckmann R
Science (2009) 326 pp. 1369-1373 [ DOI: doi:10.1126/science.1178535 Pubmed: 19933108 ]
High-resolution Cryo-EM structure of a programmed wheat germ ribosome
Resolution: 5.5 Å
EM Method: Single-particle
Fitted PDBs: 4v7e
Q-score: 0.155
Armache JP, Jarasch A, Anger AM, Villa E, Becker T, Bhushan S, Jossinet F, Habeck M, Dindar G, Franckenberg S, Marquez V, Mielke T, Thomm M, Berninghausen O, Beatrix B, Soding J, Westhof E, Wilson DN, Beckmann R
PNAS (2010) 107 pp. 19754-19759 [ Pubmed: 20974910 DOI: doi:10.1073/pnas.1010005107 ]
- Tritcum aestivum 80s ribosome (4 MDa, Complex from Triticum aestivum)
- Programmed wheat germ 80s ribosome (4 MDa, Sample)
Resolution: 6.48 Å
EM Method: Single-particle
Fitted PDBs: 2wwb
Q-score: 0.092
Becker T, Bhushan S, Jarasch A, Armache JP, Funes S, Jossinet F, Gumbart J, Mielke T, Berninghausen O, Schulten K, Westhof E, Gilmore R, Mandon EC, Beckmann R
Science (2009) 326 pp. 1369-1373 [ DOI: doi:10.1126/science.1178535 Pubmed: 19933108 ]
- Mammalian sec61 complex (70 kDa, Protein from Canis lupus familiaris)
- Actively translating wheat germ (o. sativa) 80s ribosome programmed with a nascent polypeptide chain containing a p-site trna and the type i signal anchor sequence of dpap-b (dipeptidylaminopeptidase b) bound to the mammalian (canis familiaris) sec61 complex. (4 MDa, Sample)
- Wheat germ (triticum aestivum) 80 ribosome (4 MDa, Complex)
Structure of the ribosome-bound cricket paralysis virus IRES RNA.
Resolution: 7.3 Å
EM Method: Single-particle
Fitted PDBs: 2noq
Q-score: 0.096
Schuler M, Connell SR, Lescoute A, Giesebrecht J, Dabrowski M, Schroeer B, Mielke T, Penczek PA, Westhof E, Spahn CM
Nat Struct Mol Biol (2006) 13 pp. 1092-1096 [ DOI: doi:10.1038/nsmb1177 Pubmed: 17115051 ]