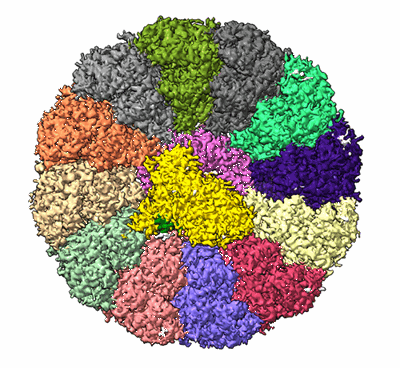
Structure of the SbCas7-11-crRNA-NTR complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8gna
Q-score: 0.544
Wang X, Yu G, Wen Y, An Q, Li X, Liao F, Lian C, Zhang K, Yin H, Wei Y, Deng Z, Zhang H
Nucleic Acids Res (2022) 50 pp. 12913-12923 [ DOI: doi:10.1093/nar/gkac1151 Pubmed: 36484100 ]
- Ternary complex (Complex from Candidatus Scalindua brodae)
- Zinc ion (65 Da, Ligand)
- Ramp superfamily protein (197 kDa, Protein from Candidatus Scalindua brodae)
- Rna (5'-r(p*gp*gp*gp*gp*cp*ap*gp*ap*ap*ap*ap*up*up*gp*gp*gp*u)-3') (14 kDa, RNA from Candidatus Scalindua brodae)
- Rna (32-mer) (10 kDa, RNA from Candidatus Scalindua brodae)
Structure of the SbCas7-11-crRNA-NTR-Csx29 complex
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8gu6
Q-score: 0.496
Wang X, Yu G, Wen Y, An Q, Li X, Liao F, Lian C, Zhang K, Yin H, Wei Y, Deng Z, Zhang H
Nucleic Acids Res (2022) 50 pp. 12913-12923 [ DOI: doi:10.1093/nar/gkac1151 Pubmed: 36484100 ]
- Rna (5'-r(p*gp*gp*gp*gp*cp*ap*gp*ap*ap*ap*ap*up*up*gp*gp*gp*u)-3') (5 kDa, RNA from Candidatus Scalindua brodae)
- Chat domain protein (82 kDa, Protein from Candidatus Scalindua brodae)
- Ternary complex (Complex from Candidatus Scalindua brodae)
- Zinc ion (65 Da, Ligand)
- Rna (33-mer) (10 kDa, RNA from Candidatus Scalindua brodae)
- Ramp superfamily protein (197 kDa, Protein from Candidatus Scalindua brodae)
Molecular architecture of the chikungunya virus replication complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7y38
Q-score: 0.539
Tan YB, Chmielewski D, Law MCY, Zhang K, He Y, Chen M, Jin J, Luo D
Sci Adv (2022) 8 pp. eadd2536-eadd2536 [ DOI: doi:10.1126/sciadv.add2536 Pubmed: 36449616 ]
- Nsp4 rdrp; docked within nsp1 central cavity (Complex from Onyong-nyong virus)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Nsp1 dodecameric ring (Complex from Chikungunya virus strain S27-African prototype)
- Protease nsp2 (89 kDa, Protein from Chikungunya virus strain S27-African prototype)
- Nsp2 helicase-protease; docked on top of viral complex with synthetic rna bound (Complex from Chikungunya virus strain S27-African prototype)
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase nsp4 (68 kDa, Protein from Onyong-nyong virus)
- Rna (5'-r(p*cp*cp*a)-3') (894 Da, RNA from in vitro transcription vector pT7-TP(deltai))
- Alphavirus replication complex (100 kDa, Complex)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Mrna-capping enzyme nsp1,affinity-tag (strepii-3xflag) (64 kDa, Protein from Chikungunya virus strain S27-African prototype)
Cryo-EM Structure of Chikungunya Virus Nonstructural Protein 1 with m7Gppp-AU
Resolution: 2.51 Å
EM Method: Single-particle
Fitted PDBs: 7fgi
Q-score: 0.627
Zhang K, Law MCY, Nguyen TM, Tan YB, Wirawan M, Law YS, Jeong LS, Luo D
Cell Rep (2022) 40 pp. 111133-111133 [ DOI: doi:10.1016/j.celrep.2022.111133 Pubmed: 35905713 ]
- Magnesium ion (24 Da, Ligand)
- Mrna-capping enzyme nsp1 (61 kDa, Protein from Chikungunya virus strain S27-African prototype)
- S-adenosyl-l-homocysteine (384 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- M7gppp-au (1 kDa, RNA from synthetic construct)
- Nonstructural protein 1 (Complex from Chikungunya virus strain S27-African prototype)
Human Drosha and DGCR8 in complex with Primary MicroRNA (MP/RNA complex) - Active state
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 6v5b
Q-score: 0.337
Partin AC, Zhang K, Jeong BC, Herrell E, Li S, Chiu W, Nam Y
Mol Cell (2020) 78 pp. 411- [ Pubmed: 32220646 DOI: doi:10.1016/j.molcel.2020.02.016 ]
- Human drosha and dgcr8 in complex with primary microrna (mp/rna complex) - active state (40 kDa, Complex from Homo sapiens)
- Ribonuclease 3 (118 kDa, Protein from Homo sapiens)
- Microprocessor complex subunit dgcr8 (60 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Pri-mir-16-2 (78-mer) (33 kDa, RNA from Homo sapiens)
Human Drosha and DGCR8 in complex with Primary MicroRNA (MP/RNA complex) - partially docked state
Resolution: 4.4 Å
EM Method: Single-particle
Fitted PDBs: 6v5c
Q-score: 0.294
Partin AC, Zhang K, Jeong BC, Herrell E, Li S, Chiu W, Nam Y
Mol Cell (2020) 78 pp. 411- [ DOI: doi:10.1016/j.molcel.2020.02.016 Pubmed: 32220646 ]
- Ribonuclease 3 (118 kDa, Protein from Homo sapiens)
- Microprocessor complex subunit dgcr8 (60 kDa, Protein from Homo sapiens)
- Pri-mir-16-2 (66-mer) (33 kDa, RNA from Homo sapiens)
- Human drosha and dgcr8 in complex with primary microrna (mp/rna complex) - partially docked state (40 kDa, Complex from Homo sapiens)
Type I-F CRISPR-Csy complex with its inhibitor AcrF9
Resolution: 2.57 Å
EM Method: Single-particle
Fitted PDBs: 6vqv
Q-score: 0.495
Zhang K, Wang S, Li S, Zhu Y, Pintilie GD, Mou TC, Schmid MF, Huang Z, Chiu W
PNAS (2020) 117 pp. 7176-7182 [ DOI: doi:10.1073/pnas.1922638117 Pubmed: 32170016 ]
- Crispr-associated protein csy1 (49 kDa, Protein from Pseudomonas aeruginosa)
- Crispr-associated protein csy3 (39 kDa, Protein from Pseudomonas aeruginosa)
- Acrf9 (7 kDa, Protein from Pseudomonas aeruginosa)
- Crrna (60-mer) (19 kDa, RNA from Pseudomonas aeruginosa)
- Crispr-associated endonuclease cas6/csy4 (21 kDa, Protein from Pseudomonas aeruginosa)
- Type i-f crispr-csy complex with its inhibitor acrf9 (350 kDa, Complex from Pseudomonas aeruginosa)
- Crispr-associated protein csy2 (36 kDa, Protein from Pseudomonas aeruginosa)
Type I-F CRISPR-Csy complex with its inhibitor AcrF8
Resolution: 3.42 Å
EM Method: Single-particle
Fitted PDBs: 6vqw
Q-score: 0.462
Zhang K, Wang S, Li S, Zhu Y, Pintilie GD, Mou TC, Schmid MF, Huang Z, Chiu W
PNAS (2020) 117 pp. 7176-7182 [ DOI: doi:10.1073/pnas.1922638117 Pubmed: 32170016 ]
- Crrna (40-mer) (19 kDa, RNA from Pseudomonas aeruginosa)
- Acrf8 (10 kDa, Protein from Pseudomonas aeruginosa)
- Crispr-associated protein csy1 (49 kDa, Protein from Pseudomonas aeruginosa)
- Crispr-associated protein csy3 (39 kDa, Protein from Pseudomonas aeruginosa)
- Type i-f crispr-associated protein csy2 (36 kDa, Protein from Pseudomonas aeruginosa)
- Type i-f crispr-csy complex with its inhibitor acrf8 (350 kDa, Complex from Pseudomonas aeruginosa)
- Crispr-associated endonuclease cas6/csy4 (21 kDa, Protein from Pseudomonas aeruginosa)
Type I-F CRISPR-Csy complex with its inhibitor AcrF6
Resolution: 3.15 Å
EM Method: Single-particle
Fitted PDBs: 6vqx
Q-score: 0.433
Zhang K, Wang S, Li S, Zhu Y, Pintilie GD, Mou TC, Schmid MF, Huang Z, Chiu W
PNAS (2020) 117 pp. 7176-7182 [ DOI: doi:10.1073/pnas.1922638117 Pubmed: 32170016 ]
- Crispr-associated endonuclease cas6/csy4 (21 kDa, Protein from Pseudomonas aeruginosa)
- Acrf6 (10 kDa, Protein from Pseudomonas aeruginosa)
- Type i-f crispr-csy complex with its inhibitor acrf6 (350 kDa, Complex from Pseudomonas aeruginosa)
- Crispr-associated protein csy1 (49 kDa, Protein from Pseudomonas aeruginosa)
- Crrna (60-mer) (19 kDa, RNA from Pseudomonas aeruginosa)
- Crispr-associated protein csy3 (39 kDa, Protein from Pseudomonas aeruginosa)
- Type i-f crispr-associated protein csy2 (36 kDa, Protein from Pseudomonas aeruginosa)
Small conformation of ssRNA-bound CRISPR_Csm complex
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 6nud
Q-score: 0.254
Guo M, Zhang K, Zhu Y, Pintilie GD, Guan X, Li S, Schmid MF, Ma Z, Chiu W, Huang Z
Cell Res (2019) 29 pp. 305-312 [ DOI: doi:10.1038/s41422-019-0151-x Pubmed: 30814678 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Small conformation of ssrna-bound crispr_csm complex (Complex from Streptococcus thermophilus)
- Crispr type iii-associated ramp protein csm4 (33 kDa, Protein from Streptococcus thermophilus)
- Target ssrna (15 kDa, RNA from Streptococcus thermophilus)
- Crispr type iii-associated ramp protein csm3 (24 kDa, Protein from Streptococcus thermophilus)
- Crispr system cms protein csm2 (14 kDa, Protein from Streptococcus thermophilus)
- Crispr system single-strand-specific deoxyribonuclease cas10/csm1 (subtype iii-a) (86 kDa, Protein from Streptococcus thermophilus)
- Crrna (22 kDa, RNA from Streptococcus thermophilus)