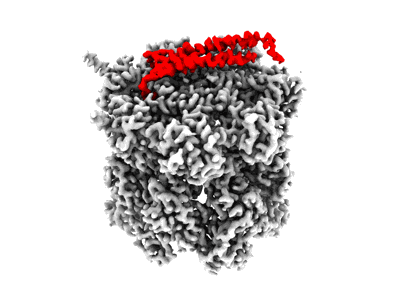
Cryo-EM map of the WT KdpFABC complex in the E1-P tight conformation, under turnover conditions
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7zre
Q-score: 0.487
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hanelt I, Paulino C
eLife (2022) 11 [ Pubmed: 36255052 DOI: doi:10.7554/eLife.80988 ]
- Potassium ion (39 Da, Ligand)
- Cardiolipin (1 kDa, Ligand)
- Potassium-transporting atpase kdpc subunit (20 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase atp-binding subunit (72 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase kdpf subunit (2 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase potassium-binding subunit (59 kDa, Protein from Escherichia coli)
- Kdpfabc (157 kDa, Complex from Escherichia coli)
Cryo-EM map of the WT KdpFABC complex in the E1_ATPearly conformation, under turnover conditions
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7zrg
Q-score: 0.465
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hanelt I, Paulino C
eLife (2022) 11 [ Pubmed: 36255052 DOI: doi:10.7554/eLife.80988 ]
- Potassium-transporting atpase kdpf subunit (2 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Kdpfabc (157 kDa, Complex from Escherichia coli)
- Potassium-transporting atpase kdpc subunit (20 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase atp-binding subunit (72 kDa, Protein from Escherichia coli)
- Cardiolipin (1 kDa, Ligand)
- Potassium ion (39 Da, Ligand)
- Potassium-transporting atpase potassium-binding subunit (59 kDa, Protein from Escherichia coli)
Cryo-EM map of the WT KdpFABC complex in the E1-P_ADP conformation, under turnover conditions
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7zrk
Q-score: 0.529
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hanelt I, Paulino C
eLife (2022) 11 [ Pubmed: 36255052 DOI: doi:10.7554/eLife.80988 ]
- Potassium-transporting atpase kdpf subunit (2 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase atp-binding subunit (72 kDa, Protein from Escherichia coli)
- Kdpfabc (157 kDa, Complex from Escherichia coli)
- Potassium-transporting atpase kdpc subunit (20 kDa, Protein from Escherichia coli)
- Cardiolipin (1 kDa, Ligand)
- Potassium ion (39 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Potassium-transporting atpase potassium-binding subunit (59 kDa, Protein from Escherichia coli)
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 7zrl
Q-score: 0.42
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hanelt I, Paulino C
eLife (2022) 11 [ DOI: doi:10.7554/eLife.80988 Pubmed: 36255052 ]
- Potassium-transporting atpase atp-binding subunit (72 kDa, Protein from Escherichia coli K-12)
- Kdpfabc (157 kDa, Complex from Escherichia coli K-12)
- Potassium-transporting atpase potassium-binding subunit (59 kDa, Protein from Escherichia coli K-12)
- Potassium-transporting atpase kdpc subunit (20 kDa, Protein from Escherichia coli K-12)
- Potassium-transporting atpase kdpf subunit (2 kDa, Protein from Escherichia coli K-12)
- Potassium ion (39 Da, Ligand)
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 7zrm
Q-score: 0.446
Silberberg JM, Stock C, Hielkema L, Corey RA, Rheinberger J, Wunnicke D, Dubach VRA, Stansfeld PJ, Hanelt I, Paulino C
eLife (2022) 11 [ Pubmed: 36255052 DOI: doi:10.7554/eLife.80988 ]
- Potassium-transporting atpase atp-binding subunit (72 kDa, Protein from Escherichia coli)
- Potassium ion (39 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Potassium-transporting atpase kdpc subunit (20 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase kdpf subunit (2 kDa, Protein from Escherichia coli)
- Potassium-transporting atpase potassium-binding subunit (59 kDa, Protein from Escherichia coli)
- Kdpfabc (157 kDa, Complex from Escherichia coli)
TWIK1 in MSP1D1 lipid nanodisc at pH 7.4
Resolution: 3.33 Å
EM Method: Single-particle
Fitted PDBs: 7sk0
Q-score: 0.357
Nat Commun (2022) 13 pp. 3232-3232 [ Pubmed: 35680900 DOI: doi:10.1038/s41467-022-30853-z ]
- N-octane (114 Da, Ligand)
- Potassium channel subfamily k member 1 (38 kDa, Protein from Rattus norvegicus)
- Potassium ion (39 Da, Ligand)
- Twik1 in msp1d1 lipid nanodisc at ph 7.4 (Complex from Rattus norvegicus)
Akirin2 bound human proteasome
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 7nht
Q-score: 0.547
de Almeida M, Hinterndorfer M, Brunner H, Grishkovskaya I, Singh K, Schleiffer A, Jude J, Deswal S, Kalis R, Vunjak M, Lendl T, Imre R, Roitinger E, Neumann T, Kandolf S, Schutzbier M, Mechtler K, Versteeg GA, Haselbach D, Zuber J
Nature (2021) 599 pp. 491-496 [ DOI: doi:10.1038/s41586-021-04035-8 Pubmed: 34711951 ]
- Potassium ion (39 Da, Ligand)
- Proteasome subunit beta type-5 (28 kDa, Protein from Human)
- Proteasome subunit beta type-4 (29 kDa, Protein from Human)
- Proteasome subunit beta type-6 (25 kDa, Protein from Human)
- Proteasome subunit alpha type-3 (28 kDa, Protein from Human)
- Akirin2 bound to 20s proteasome (780 kDa, Complex)
- Akirin2 (Complex from Homo sapiens)
- 20s proteasome (Complex from Homo sapiens)
- Proteasome subunit alpha type-5 (26 kDa, Protein from Human)
- Proteasome subunit beta type-2 (22 kDa, Protein from Human)
- Proteasome subunit alpha type-1 (29 kDa, Protein from Human)
- Proteasome subunit alpha type-6 (27 kDa, Protein from Human)
- Proteasome subunit beta type-3 (22 kDa, Protein from Human)
- Proteasome subunit alpha type-2 (25 kDa, Protein from Human)
- Proteasome subunit alpha type-7 (27 kDa, Protein from Human)
- Proteasome subunit beta type-1 (26 kDa, Protein from Human)
- Akirin-2 (22 kDa, Protein from Homo sapiens)
- Proteasome subunit alpha type-4 (29 kDa, Protein from Human)
- Proteasome subunit beta type-7 (30 kDa, Protein from Human)
Cryo-EM structure of the potassium-chloride cotransporter KCC4 in lipid nanodiscs
Resolution: 3.65 Å
EM Method: Single-particle
Fitted PDBs: 6ukn
Q-score: 0.456
eLife (2020) 9 [ Pubmed: 32286222 DOI: doi:10.7554/eLife.52505 ]
- Chloride ion (35 Da, Ligand)
- Potassium ion (39 Da, Ligand)
- Solute carrier family 12 member 7 (120 kDa, Protein from Mus musculus)
- Kcc4 in lipid nanodiscs (Cell from Mus musculus)
Cryo-EM structure of human KATP bound to ATP and ADP in quatrefoil form
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 6c3o
Q-score: 0.379
eLife (2017) 6 [ Pubmed: 29286281 DOI: doi:10.7554/eLife.32481 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Quatrefoil form of human katp in complex with atp and adp (880 MDa, Complex from Homo sapiens)
- Potassium ion (39 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Atp-sensitive inward rectifier potassium channel 11 (45 kDa, Protein from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Atp-binding cassette sub-family c member 8 (177 kDa, Protein from Homo sapiens)