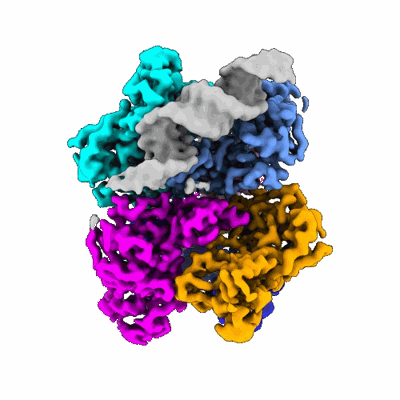
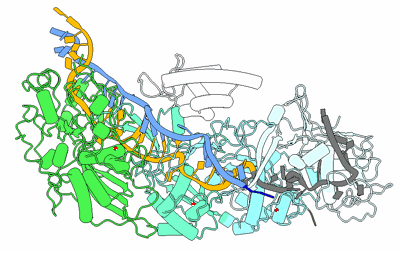
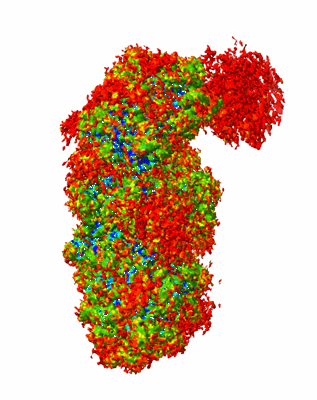
SpCas9-MMLV RT-pegRNA-target DNA complex (initiation)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8wut
Q-score: 0.364
Shuto Y, Nakagawa R, Zhu S, Hoki M, Omura SN, Hirano H, Itoh Y, Zhang F, Nureki O
Nature (2024) [ DOI: doi:10.1038/s41586-024-07497-8 Pubmed: 38811740 ]
- Crispr-associated endonuclease cas9/csn1 (158 kDa, Protein from Streptococcus pyogenes serotype M1)
- Dna/rna (35-mer) (10 kDa, Macromolecule from Streptococcus pyogenes)
- Dna (5'-d(*tp*gp*ap*tp*gp*gp*cp*ap*gp*ap*gp*tp*ap*cp*tp*ap*g)-3') (5 kDa, DNA from Streptococcus pyogenes)
- Gag-pol polyprotein (55 kDa, Protein from Moloney murine leukemia virus)
- Spcas9-mmlv rt-pegrna-target dna complex (Complex from Streptococcus pyogenes)
- Dna (51-mer) (15 kDa, DNA from Streptococcus pyogenes)
- Rna (115-mer) (37 kDa, RNA from Streptococcus pyogenes)
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, consensus form
Resolution: 2.67 Å
EM Method: Single-particle
Fitted PDBs: 8ud3
Q-score: 0.605
Ito F, Yang H, Zhou ZH, Chen XS
Protein Cell (2024) [ Pubmed: 38613382 DOI: doi:10.1093/procel/pwae009 ]
- Homohexamer of nsp15 in complex with poly(a/u) rna (262 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (35-mer) (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 15 (40 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (35-mer) (11 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 1
Resolution: 3.25 Å
EM Method: Single-particle
Fitted PDBs: 8ud4
Q-score: 0.511
Ito F, Yang H, Zhou ZH, Chen XS
Protein Cell (2024) [ Pubmed: 38613382 DOI: doi:10.1093/procel/pwae009 ]
- Non-structural protein 15 (40 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (35-mer) (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (35-mer) (11 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Homohexamer of nsp15 in complex with rna duplex (262 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 Nsp15 bound to poly(A/U) RNA, state 2
Resolution: 3.13 Å
EM Method: Single-particle
Fitted PDBs: 8ud5
Q-score: 0.541
Ito F, Yang H, Zhou ZH, Chen XS
Protein Cell (2024) [ Pubmed: 38613382 DOI: doi:10.1093/procel/pwae009 ]
- Rna (35-mer) (11 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Homohexamer of nsp15 in complex with rna duplex (262 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (35-mer) (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 15 (40 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
respiratory syncytial virus nucleocapsid-like assembly
Resolution: 3.96 Å
EM Method: Single-particle
Fitted PDBs: 8iuo
Q-score: 0.364
Wang Y, Zhang C, Luo Y, Ling X, Luo B, Jia G, Su D, Dong H, Su Z
Signal Transduct Target Ther (2023) 8 pp. 323-323 [ Pubmed: 37607909 DOI: doi:10.1038/s41392-023-01602-5 ]
- Nucleoprotein (40 kDa, Protein from Human respiratory syncytial virus A)
- Rna (35-mer) (10 kDa, RNA from Homo sapiens)
- Respiratory syncytial virus nucleocapsid-like assembly (Complex from Human respiratory syncytial virus A)
gRAMP non-matching PFS-with Mg
Resolution: 3.62 Å
EM Method: Single-particle
Fitted PDBs: 8d9i
Q-score: 0.423
Hu C, van Beljouw SPB, Nam KH, Schuler G, Ding F, Cui Y, Rodriguez-Molina A, Haagsma AC, Valk M, Pabst M, Brouns SJJ, Ke A
Science (2022) 377 pp. 1278-1285 [ Pubmed: 36007061 DOI: doi:10.1126/science.add5064 ]
- Ramp superfamily protein (143 kDa, Protein from Candidatus Scalindua brodae)
- Rna (5'-r(p*up*cp*cp*gp*gp*gp*gp*cp*ap*gp*ap*ap*ap*ap*up*up*gp*gp*a)-3') (6 kDa, RNA from Candidatus Scalindua brodae)
- Gramp-non matching pfs-with mg (Complex from Candidatus Scalindua brodae)
- Zinc ion (65 Da, Ligand)
- Rna (35-mer) (11 kDa, RNA from Candidatus Scalindua brodae)
Structure of the type III-E CRISPR-Cas effector gRAMP
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 7xso
Q-score: 0.553
Liu X, Zhang L, Wang H, Xiu Y, Huang L, Gao Z, Li N, Li F, Xiong W, Gao T, Zhang Y, Yang M, Feng Y
Mol Cell (2022) 82 pp. 4503-4518.e8 [ DOI: doi:10.1016/j.molcel.2022.10.007 Pubmed: 36306795 ]
- Gramp (Complex from Candidatus Scalindua brodae)
- Rna (35-mer) (23 kDa, RNA from Candidatus Scalindua brodae)
- Ramp superfamily protein (197 kDa, Protein from Candidatus Scalindua brodae)
- Zinc ion (65 Da, Ligand)
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 7uo4
Q-score: 0.457
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ DOI: doi:10.1038/s41586-022-05664-3 Pubmed: 36725929 ]
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Sars-cov-2 replication-transcription complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Water (18 Da, Ligand)
- [[(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate (531 Da, Ligand)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Resolution: 3.09 Å
EM Method: Single-particle
Fitted PDBs: 7uo7
Q-score: 0.451
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ Pubmed: 36725929 DOI: doi:10.1038/s41586-022-05664-3 ]
- Magnesium ion (24 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 replication-transcription complex + atp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Adenosine-5'-triphosphate (507 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state.
Resolution: 3.13 Å
EM Method: Single-particle
Fitted PDBs: 7uo9
Q-score: 0.487
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ Pubmed: 36725929 DOI: doi:10.1038/s41586-022-05664-3 ]
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 replication-transcription complex + utp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Uridine 5'-triphosphate (484 Da, Ligand)
- Water (18 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)