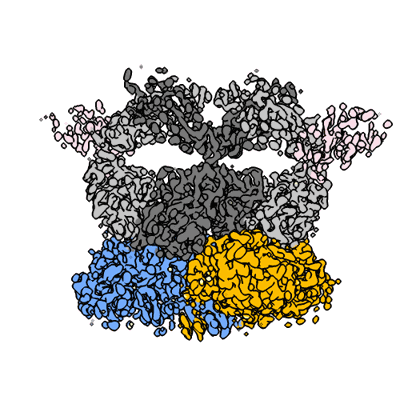
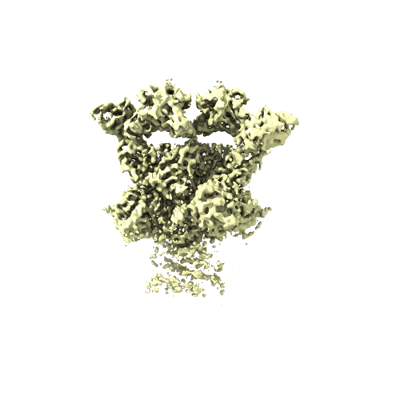
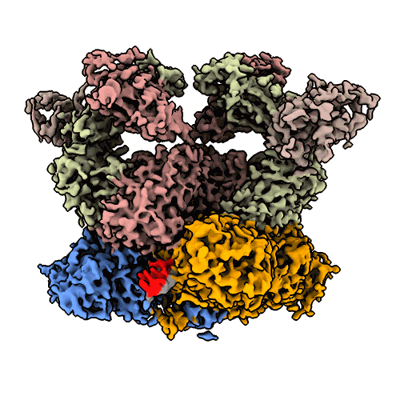
ZIKV rsNS1 in complex with Fab AA12 and anti-fab nanobody
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8wnp
Q-score: 0.428
Chew BLA, Ngoh AQ, Phoo WW, Weng MJG, Sheng HJ, Chan KWK, Tan EYJ, Gelbart T, Xu C, Tan GS, Vasudevan SG, Luo D
Npj Viruses (2024) 2 pp. 14- [ DOI: doi:10.1038/s44298-024-00024-6 ]
- Vh chains (23 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Vl chains (23 kDa, Protein from Homo sapiens)
- Anti-fab nanobody (13 kDa, Protein from Lama glama)
- Zikv rsns1 in complex with fab aa12 and anti-fab nanobody (Complex from Zika virus)
- Non-structural protein 1 (40 kDa, Protein from Zika virus)
ZIKV rsNS1 in complex with Fab GB5 and anti-fab nanobody
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 8wnu
Q-score: 0.331
Chew BLA, Ngoh AQ, Phoo WW, Weng MJG, Sheng HJ, Chan KWK, Tan EYJ, Gelbart T, Xu C, Tan GS, Vasudevan SG, Luo D
Npj Viruses (2024) 2 pp. 14- [ DOI: doi:10.1038/s44298-024-00024-6 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Anti-fab nanobody (13 kDa, Protein from Lama glama)
- Vl (23 kDa, Protein from Homo sapiens)
- Non-structural protein 1 (40 kDa, Protein from Zika virus)
- Zikv rsns1 in complex with fab gb5 and anti-fab nanobody (Complex from Zika virus)
- Vh (23 kDa, Protein from Homo sapiens)
ZIKV rsNS1 in complex with Fab EB9 and anti-fab nanobody
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8wo4
Q-score: 0.416
Chew BLA, Ngoh AQ, Phoo WW, Weng MJG, Sheng HJ, Chan KWK, Tan EYJ, Gelbart T, Xu C, Tan GS, Vasudevan SG, Luo D
Npj Viruses (2024) 2 pp. 14- [ DOI: doi:10.1038/s44298-024-00024-6 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Non-structural protein 1 (41 kDa, Protein from Zika virus)
- Zikv rsns1 in complex with fab aa12 and anti-fab nanobody (Complex from Homo sapiens)
- Vl (23 kDa, Protein from Homo sapiens)
- Vh (24 kDa, Protein from Homo sapiens)
- Anti-fab nanobody (15 kDa, Protein from Lama glama)
Cryo-EM structure of the GI.4 Chiba VLP complexed with the CV-1A1 Fv-clasp
Resolution: 3.04 Å
EM Method: Single-particle
Fitted PDBs: 8jg5
Q-score: 0.376
Kimura-Someya T, Katsura K, Kato-Murayama M, Hosaka T, Uchikubo-Kamo T, Ihara K, Hanada K, Sato S, Murayama K, Kataoka M, Shirouzu M, Someya Y
J Virol (2024) 98 pp. e0019724-e0019724 [ Pubmed: 38593321 DOI: doi:10.1128/jvi.00197-24 ]
- Vh,sarah (19 kDa, Protein from Homo sapiens)
- Vl,sarah (18 kDa, Protein from Homo sapiens)
- Viral protein 1 with its antibody fragments (Complex from Chiba virus)
- Vp1 (58 kDa, Protein from Chiba virus)
SARS-CoV-2 Omicron 1-RBD up Spike trimer complexed with three XG005 molecules
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 7ycy
Q-score: 0.26
To Be Published
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Xg005-vl (11 kDa, Protein from Homo sapiens)
- Xg005-vh (13 kDa, Protein from Homo sapiens)
SARS-CoV-2 Omicron 2-RBD up Spike trimer complexed with three XG005 molecules
Resolution: 3.24 Å
EM Method: Single-particle
Fitted PDBs: 7ycz
Q-score: 0.352
To Be Published
- Xg005-vl (21 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Xg005-vh (23 kDa, Protein from Homo sapiens)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 Omicron 1-RBD up spike trimer complexed with two XG005 Fab
Resolution: 3.58 Å
EM Method: Single-particle
Fitted PDBs: 7yd0
Q-score: 0.267
To Be Published
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Xg005-vh (23 kDa, Protein from Homo sapiens)
- Xg005-vl (21 kDa, Protein from Homo sapiens)
Structure of X18 UFO protomer in complex with F6 Fab VHVL domain
Resolution: 4.05 Å
EM Method: Single-particle
Fitted PDBs: 8gp5
Q-score: 0.422
Niu J, Wang Q, Zhao W, Meng B, Xu Y, Zhang X, Feng Y, Qi Q, Hao Y, Zhang X, Liu Y, Xiang J, Shao Y, Yang B
Nat Commun (2023) 14 pp. 4676-4676 [ Pubmed: 37542068 DOI: doi:10.1038/s41467-023-40321-x ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- X18 ufo env protomer (69 kDa, Protein from Homo sapiens)
- F6 fab vh domain (25 kDa, Protein from Homo sapiens)
- F6 fab vl domain (24 kDa, Protein from Homo sapiens)
- X18 ufo protomer in complex with f6 vhvl domain (Complex from Homo sapiens)
Local refinement of SARS-CoV-2 Omicron S trimer complexed with XG005
Resolution: 2.99 Å
EM Method: Single-particle
Fitted PDBs: 7yd1
Q-score: 0.432
To Be Published
- Xg005-vl (11 kDa, Protein from Homo sapiens)
- Xg005-vh (13 kDa, Protein from Homo sapiens)
- Spike protein s1 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Omicron rbd with xg005 (Complex from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM Structure of HPIV3 prefusion F trimer in complex with 3x1 Fab
Resolution: 3.62 Å
EM Method: Single-particle
Fitted PDBs: 8dg8
Q-score: 0.39
Caban M, Rodarte JV, Bibby M, Gray MD, Taylor JJ, Pancera M, Boonyaratanakornkit J
Nat Commun (2023) 14 pp. 798-798 [ Pubmed: 36781872 DOI: doi:10.1038/s41467-023-36459-3 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- 3x1 monoclonal antibody, vh region (13 kDa, Protein from Homo sapiens)
- Complex of one 3x1 fab bound to the hpiv3 pref trimer (339 kDa, Complex from Homo sapiens)
- 3x1 monoclonal antibody, vl region (11 kDa, Protein from Homo sapiens)
- Fusion glycoprotein f0 (54 kDa, Protein from Human parainfluenza 3 virus (strain NIH 47885))