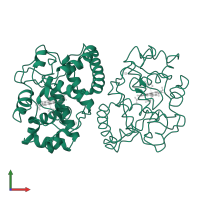Assemblies
Assembly Name:
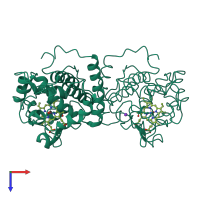
L-ascorbate peroxidase, cytosolic
Multimeric state:
homo dimer
Accessible surface area:
19508.51 Å2
Buried surface area:
4250.97 Å2
Dissociation area:
865.35
Å2
Dissociation energy (ΔGdiss):
-6.13
kcal/mol
Dissociation entropy (TΔSdiss):
12.94
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-155749
Assembly Name:
L-ascorbate peroxidase, cytosolic
Multimeric state:
homo dimer
Accessible surface area:
19777.67 Å2
Buried surface area:
4197.9 Å2
Dissociation area:
836.8
Å2
Dissociation energy (ΔGdiss):
-5.55
kcal/mol
Dissociation entropy (TΔSdiss):
12.93
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-155749
Macromolecules
Chains: A, B, C, D
Length: 249 amino acids
Theoretical weight: 27.1 KDa
Source organism: Pisum sativum
Expression system: Escherichia coli
UniProt:
Pfam: Peroxidase
InterPro:
SCOP: CCP-like
Length: 249 amino acids
Theoretical weight: 27.1 KDa
Source organism: Pisum sativum
Expression system: Escherichia coli
UniProt:
- Canonical:
 P48534 (Residues: 2-250; Coverage: 100%)
P48534 (Residues: 2-250; Coverage: 100%)
Pfam: Peroxidase
InterPro:
- Haem peroxidase superfamily
- Heme-binding peroxidase Ccp1-like
- Class I peroxidase
- Haem peroxidase
- Peroxidase, active site
- Peroxidases heam-ligand binding site
SCOP: CCP-like