Assemblies
Assembly Name:
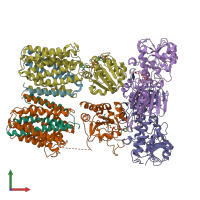
proton-translocating NAD(P)(+) transhydrogenase and NAD(P) transhydrogenase subunit beta
Multimeric state:
hetero hexamer
Accessible surface area:
68091.41 Å2
Buried surface area:
30849.43 Å2
Dissociation area:
2,282.62
Å2
Dissociation energy (ΔGdiss):
19.69
kcal/mol
Dissociation entropy (TΔSdiss):
15.63
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-180813
Macromolecules
Chains: A, C
Length: 100 amino acids
Theoretical weight: 10.65 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
Pfam: 4TM region of pyridine nucleotide transhydrogenase, mitoch
InterPro: NAD(P) transhydrogenase, alpha subunit, C-terminal
Length: 100 amino acids
Theoretical weight: 10.65 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q72GR9 (Residues: 1-100; Coverage: 100%)
Q72GR9 (Residues: 1-100; Coverage: 100%)
Pfam: 4TM region of pyridine nucleotide transhydrogenase, mitoch
InterPro: NAD(P) transhydrogenase, alpha subunit, C-terminal
Chains: B, D
Length: 450 amino acids
Theoretical weight: 47.37 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
Pfam: NAD(P) transhydrogenase beta subunit
InterPro:
Length: 450 amino acids
Theoretical weight: 47.37 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q72GS0 (Residues: 1-450; Coverage: 100%)
Q72GS0 (Residues: 1-450; Coverage: 100%)
Pfam: NAD(P) transhydrogenase beta subunit
InterPro:
Chains: E, F
Length: 384 amino acids
Theoretical weight: 41.26 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 384 amino acids
Theoretical weight: 41.26 KDa
Source organism: Thermus thermophilus HB27
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q72GR8 (Residues: 1-375; Coverage: 100%)
Q72GR8 (Residues: 1-375; Coverage: 100%)
Pfam:
InterPro: