Assemblies
Assembly Name:
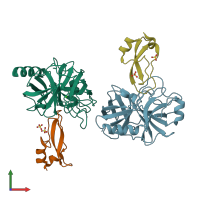
Pancreatic trypsin inhibitor and Serine protease 1
Multimeric state:
hetero dimer
Accessible surface area:
11942.59 Å2
Buried surface area:
1807.55 Å2
Dissociation area:
717.71
Å2
Dissociation energy (ΔGdiss):
3.68
kcal/mol
Dissociation entropy (TΔSdiss):
10.46
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-133752
Assembly Name:
Pancreatic trypsin inhibitor and Serine protease 1
Multimeric state:
hetero dimer
Accessible surface area:
12039.41 Å2
Buried surface area:
1891.75 Å2
Dissociation area:
716.12
Å2
Dissociation energy (ΔGdiss):
4.97
kcal/mol
Dissociation entropy (TΔSdiss):
10.45
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-133752
Macromolecules
Chains: A, B
Length: 224 amino acids
Theoretical weight: 24.09 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Trypsin
InterPro:
Length: 224 amino acids
Theoretical weight: 24.09 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P07477 (Residues: 24-247; Coverage: 97%)
P07477 (Residues: 24-247; Coverage: 97%)
Pfam: Trypsin
InterPro:
- Serine proteases, trypsin domain
- Peptidase S1, PA clan
- Peptidase S1, PA clan, chymotrypsin-like fold
- Peptidase S1A, chymotrypsin family
- Serine proteases, trypsin family, histidine active site
Chains: C, I
Length: 55 amino acids
Theoretical weight: 6.34 KDa
Source organism: Bos taurus
Expression system: Komagataella pastoris
UniProt:
InterPro:
Length: 55 amino acids
Theoretical weight: 6.34 KDa
Source organism: Bos taurus
Expression system: Komagataella pastoris
UniProt:
- Canonical:
 P00974 (Residues: 36-90; Coverage: 70%)
P00974 (Residues: 36-90; Coverage: 70%)
InterPro:
- Pancreatic trypsin inhibitor Kunitz domain superfamily
- Pancreatic trypsin inhibitor Kunitz domain
- Proteinase inhibitor I2, Kunitz, conserved site