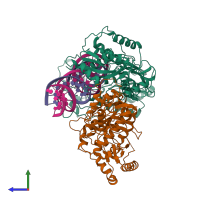
Assemblies
Assembly Name:
Protease and RNA
Multimeric state:
hetero tetramer
Accessible surface area:
54678.51 Å2
Buried surface area:
9917.08 Å2
Dissociation area:
1,202.46
Å2
Dissociation energy (ΔGdiss):
7.2
kcal/mol
Dissociation entropy (TΔSdiss):
13.65
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-249588
Macromolecules
Chain: A
Length: 562 amino acids
Theoretical weight: 64.71 KDa
Source organism: Human immunodeficiency virus type 1 BH10
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 562 amino acids
Theoretical weight: 64.71 KDa
Source organism: Human immunodeficiency virus type 1 BH10
Expression system: Escherichia coli
UniProt:
- Canonical:
 P03366 (Residues: 600-1159; Coverage: 39%)
P03366 (Residues: 600-1159; Coverage: 39%)
Pfam:
- Reverse transcriptase (RNA-dependent DNA polymerase)
- Reverse transcriptase thumb domain
- Reverse transcriptase connection domain
- RNase H
Chain: B
Length: 442 amino acids
Theoretical weight: 51.59 KDa
Source organism: Human immunodeficiency virus 1
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 442 amino acids
Theoretical weight: 51.59 KDa
Source organism: Human immunodeficiency virus 1
Expression system: Escherichia coli
UniProt:
- Canonical:
 P04585 (Residues: 588-1027; Coverage: 31%)
P04585 (Residues: 588-1027; Coverage: 31%)
Pfam:
- Reverse transcriptase (RNA-dependent DNA polymerase)
- Reverse transcriptase thumb domain
- Reverse transcriptase connection domain
Name:
HIV-1 viral RNA genome fragment
Representative chains: C
Length: 101 nucleotides
Theoretical weight: 32.62 KDa
Rfam: HIV primer binding site (PBS)
Representative chains: C
Length: 101 nucleotides
Theoretical weight: 32.62 KDa
Rfam: HIV primer binding site (PBS)
Name:
tRNA lysine 3
Representative chains: D
Length: 77 nucleotides
Theoretical weight: 24.72 KDa
Rfam: tRNA
Representative chains: D
Length: 77 nucleotides
Theoretical weight: 24.72 KDa
Rfam: tRNA