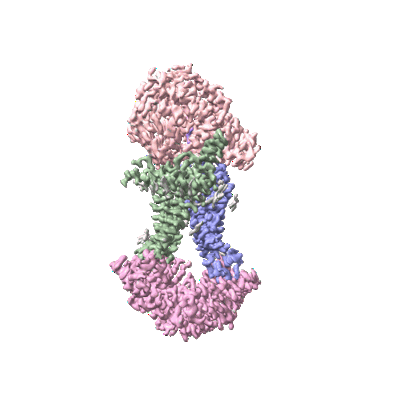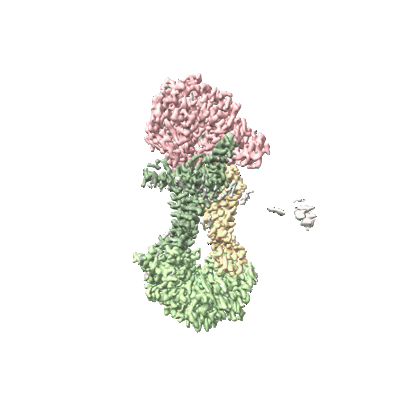Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-translocation state
Resolution: 3.25 Å
EM Method: Single-particle
Fitted PDBs: 8j5q
Q-score: 0.519
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-translocation state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Endogenous oligopeptide (783 Da, Protein from Mycolicibacterium smegmatis MC2 155)
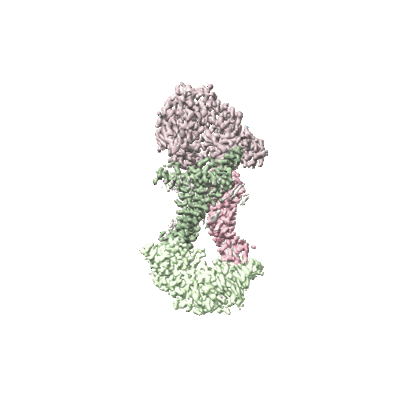
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the resting state
Resolution: 3.28 Å
EM Method: Single-particle
Fitted PDBs: 8j5r
Q-score: 0.534
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the resting state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-catalytic intermediate state
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8j5s
Q-score: 0.566
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Endogenous oligopeptide (698 Da, Protein from Mycolicibacterium smegmatis MC2 155)
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Magnesium ion (24 Da, Ligand)
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
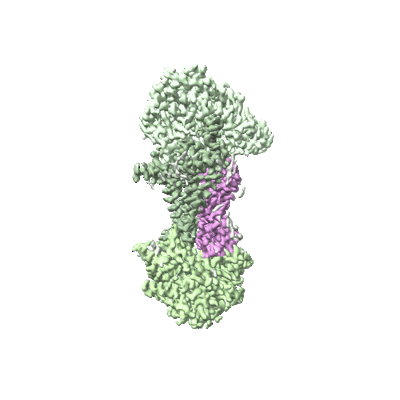
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the catalytic intermediate state
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8j5t
Q-score: 0.572
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Magnesium ion (24 Da, Ligand)
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8gwk
Q-score: 0.423
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- [(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methyl dihydrogen phosphate (371 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_rmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 E-RTC bound with MMP-nsp9 and GMPPNP
Resolution: 2.64 Å
EM Method: Single-particle
Fitted PDBs: 8gwm
Q-score: 0.41
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- 2'-deoxy-2'-fluoro-2'-methyluridine 5'-(trihydrogen diphosphate) (420 Da, Ligand)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (26-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (25-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_mmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.31 Å
EM Method: Single-particle
Fitted PDBs: 8gw1
Q-score: 0.332
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_ump-nsp9 (350 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Manganese (ii) ion (54 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Uridine-5'-monophosphate (324 Da, Ligand)
- Water (18 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Replicase polyprotein 1ab (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna (25-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of Mycobacterium tuberculosis LpqY-SugABC in complex with trehalose
Resolution: 3.02 Å
EM Method: Single-particle
Fitted PDBs: 8ja7
Q-score: 0.428
Liang J, Liu F, Xu P, Shangguan W, Hu T, Wang S, Yang X, Xiong Z, Yang X, Guddat LW, Yu B, Rao Z, Zhang B
PNAS (2023) 120 pp. e2307625120-e2307625120 [ Pubmed: 37603751 DOI: doi:10.1073/pnas.2307625120 ]
- Trehalose transport system permease protein sugb (29 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Trehalose-binding lipoprotein lpqy (49 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Trehalose transport system permease protein suga (33 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Lpqy-sugabc (Complex from Mycobacterium tuberculosis H37Rv)
- Trehalose import atp-binding protein sugc (42 kDa, Protein from Mycobacterium tuberculosis H37Rv)
Resolution: 4.29 Å
EM Method: Single-particle
Fitted PDBs: 8hpl
Q-score: 0.292
Liang J, Yang X, Hu T, Gao Y, Yang Q, Yang H, Peng W, Zhou X, Guddat LW, Zhang B, Rao Z, Liu F
Structure (2023) 31 pp. 1158-1165.e3 [ Pubmed: 37619560 DOI: doi:10.1016/j.str.2023.07.014 ]
- Abc transporter, atp-binding protein sugc (43 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Abc transporter, permease protein sugb (29 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Trehalose-specific importer lpqy-sugabc complex (Complex from Mycolicibacterium smegmatis MC2 155)
- Abc sugar transporter, permease component (32 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Bacterial extracellular solute-binding protein (50 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
Resolution: 3.82 Å
EM Method: Single-particle
Fitted PDBs: 8hpm
Q-score: 0.347
Liang J, Yang X, Hu T, Gao Y, Yang Q, Yang H, Peng W, Zhou X, Guddat LW, Zhang B, Rao Z, Liu F
Structure (2023) 31 pp. 1158-1165.e3 [ Pubmed: 37619560 DOI: doi:10.1016/j.str.2023.07.014 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Bacterial extracellular solute-binding protein (50 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Abc transporter, permease protein sugb (29 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Abc sugar transporter, permease component (32 kDa, Protein from Mycolicibacterium smegmatis MC2 155)
- Trehalose-specific importer lpqy-sugabc complex (Complex from Mycolicibacterium smegmatis MC2 155)
- Abc transporter, atp-binding protein sugc (43 kDa, Protein from Mycolicibacterium smegmatis MC2 155)