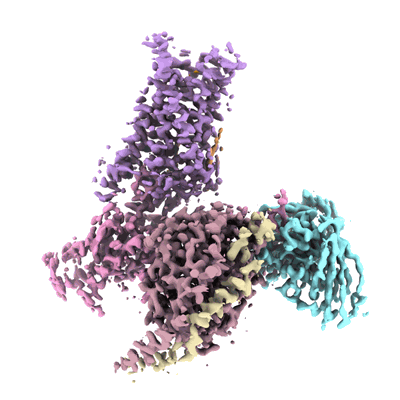
RBD of SARS-CoV2 spike protein with ACE2 decoy
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8jje
Q-score: 0.488
Urano E, Itoh Y, Suzuki T, Sasaki T, Kishikawa JI, Akamatsu K, Higuchi Y, Sakai Y, Okamura T, Mitoma S, Sugihara F, Takada A, Kimura M, Nakao S, Hirose M, Sasaki T, Koketsu R, Tsuji S, Yanagida S, Shioda T, Hara E, Matoba S, Matsuura Y, Kanda Y, Arase H, Okada M, Takagi J, Kato T, Hoshino A, Yasutomi Y, Saito A, Okamoto T
Sci Transl Med (2023) 15 pp. eadi2623-eadi2623 [ DOI: doi:10.1126/scitranslmed.adi2623 Pubmed: 37647387 ]
- Sars-cov2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Rbd of sars-cov2 spike protein with ace2 decoy. (460 kDa, Complex)
- Spike glycoprotein (138 kDa, Protein from synthetic construct)
- Ace2 (Complex from Homo sapiens)
- Sulfate ion (96 Da, Ligand)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Angiotensin-converting enzyme 2 (73 kDa, Protein from Homo sapiens)
Cryo-EM structure of the histamine-bound histamine H4 receptor and Gq complex
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7yfc
Q-score: 0.539
Im D, Kishikawa JI, Shiimura Y, Hisano H, Ito A, Fujita-Fujiharu Y, Sugita Y, Noda T, Kato T, Asada H, Iwata S
Nat Commun (2023) 14 pp. 6538-6538 [ Pubmed: 37863901 DOI: doi:10.1038/s41467-023-42260-z ]
- Nb35 (14 kDa, Protein from Lama glama)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (Complex from Lama glama)
- Nb35 (Complex from Homo sapiens)
- Scfv16 (27 kDa, Protein from Mus musculus)
- Histamine-bound histamine h4 receptor in complex with gq (190 kDa, Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (Complex from Mus musculus)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (42 kDa, Protein from Homo sapiens)
- Histamine h4 receptor (62 kDa, Protein from Homo sapiens)
- Engineered g-alpha-q (Complex from Homo sapiens)
- Histamine h4 receptor (Complex)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (7 kDa, Protein from Homo sapiens)
- Engineered g-alpha-q (41 kDa, Protein from Homo sapiens)
- Cholesterol (386 Da, Ligand)
- Histamine (111 Da, Ligand)
- Scfv16 (Complex from Homo sapiens)
Cryo-EM structure of the imetit-bound histamine H4 receptor and Gq complex
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7yfd
Q-score: 0.552
Im D, Kishikawa JI, Shiimura Y, Hisano H, Ito A, Fujita-Fujiharu Y, Sugita Y, Noda T, Kato T, Asada H, Iwata S
Nat Commun (2023) 14 pp. 6538-6538 [ Pubmed: 37863901 DOI: doi:10.1038/s41467-023-42260-z ]
- Imetit bound histamine h4 receptor in complex with gq (190 kDa, Complex from Homo sapiens)
- Nb35 (14 kDa, Protein from Lama glama)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (Complex from Lama glama)
- Nb35 (Complex)
- Scfv16 (27 kDa, Protein from Mus musculus)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (Complex from Mus musculus)
- Histamine h4 receptor (Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (42 kDa, Protein from Homo sapiens)
- Engineered g-alpha-q (Complex from Homo sapiens)
- Histamine h4 receptor (62 kDa, Protein from Homo sapiens)
- 2-(1~{h}-imidazol-5-yl)ethyl carbamimidothioate (170 Da, Ligand)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (7 kDa, Protein from Homo sapiens)
- Engineered g-alpha-q (41 kDa, Protein from Homo sapiens)
- Cholesterol (386 Da, Ligand)
- Scfv16 (Complex from Homo sapiens)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 1
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk3
Q-score: 0.567
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ Pubmed: 35882843 DOI: doi:10.1038/s41467-022-31718-1 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 1 (220 kDa, Complex from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Calcium ion (40 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Riboflavin (376 Da, Ligand)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 2
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk4
Q-score: 0.562
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ Pubmed: 35882843 DOI: doi:10.1038/s41467-022-31718-1 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 2 (220 kDa, Complex from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Calcium ion (40 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Riboflavin (376 Da, Ligand)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 3
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk5
Q-score: 0.546
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ Pubmed: 35882843 DOI: doi:10.1038/s41467-022-31718-1 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Calcium ion (40 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Riboflavin (376 Da, Ligand)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 3 (220 kDa, Complex from Vibrio cholerae O395)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7xk6
Q-score: 0.575
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ Pubmed: 35882843 DOI: doi:10.1038/s41467-022-31718-1 ]
- Flavin mononucleotide (456 Da, Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, with aurachin d-42 (220 kDa, Complex from Vibrio cholerae O395)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Aurachin d (363 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Water (18 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Riboflavin (376 Da, Ligand)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 7xk7
Q-score: 0.584
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ Pubmed: 35882843 DOI: doi:10.1038/s41467-022-31718-1 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Korormicin (433 Da, Ligand)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, with korormicin (220 kDa, Complex from Vibrio cholerae O395)
- Water (18 Da, Ligand)
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Calcium ion (40 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Riboflavin (376 Da, Ligand)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
Cryo-EM structure of MrgD-Gi complex with beta-alanine
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7y12
Q-score: 0.519
Suzuki S, Iida M, Hiroaki Y, Tanaka K, Kawamoto A, Kato T, Oshima A
Commun Biol (2022) 5 pp. 707-707 [ Pubmed: 35840655 DOI: doi:10.1038/s42003-022-03668-3 ]
- Scfv16 (31 kDa, Protein from synthetic construct)
- Guanine nucleotide-binding protein g(i) subunit alpha-1 (40 kDa, Protein from Homo sapiens)
- Beta-alanine bound mrgd-gi complex with the scfv16 (Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (7 kDa, Protein from Mus musculus)
- Beta-alanine (89 Da, Ligand)
- Subunit beta-1,subunit gamma-2 (Complex from synthetic construct)
- Palmitic acid (256 Da, Ligand)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (39 kDa, Protein from Mus musculus)
- Soluble cytochrome b562,mas-related g-protein coupled receptor member d (50 kDa, Protein from Homo sapiens)
- G(i) subunit alpha-1,mas-related g-protein coupled receptor member d (Complex from Mus musculus)
- Scfv16 (Complex)
Cryo-EM structure of apo-state MrgD-Gi complex (local)
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7y13
Q-score: 0.512
Suzuki S, Iida M, Hiroaki Y, Tanaka K, Kawamoto A, Kato T, Oshima A
Commun Biol (2022) 5 pp. 707-707 [ DOI: doi:10.1038/s42003-022-03668-3 Pubmed: 35840655 ]
- Palmitic acid (256 Da, Ligand)
- Soluble cytochrome b562,mas-related g-protein coupled receptor member d (Complex from Homo sapiens)
- Water (18 Da, Ligand)
- Soluble cytochrome b562,mas-related g-protein coupled receptor member d (50 kDa, Protein from Homo sapiens)