3DEM History and Genealogy 1968-2011
Welcome to the Web Page on the History of 3-Dimensional Electron Microscopy in Biology
Since the inception of the field of 3-Dimensional Electron Microscopy in Biology in 1968 there has been remarkable growth in the number of labs and scientists active in the field of 3DEM.
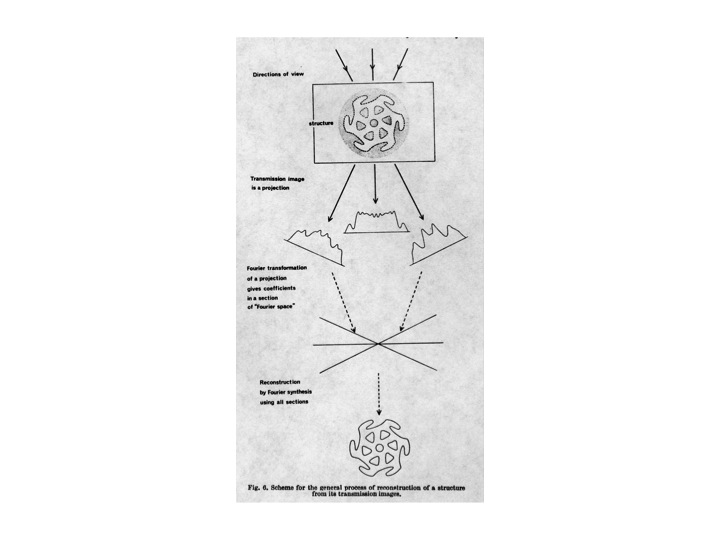
The aim of this website is to provide links to some of the original papers which spawned the field and reviews which have chronicled the subsequent development of the field. Several of these resources are published personal accountings by colleagues who were central in the field, and also narratives written especially for this project.
An attempt has been made to present a genealogy reflecting the original groups in the field and to show how the field has propagated from the few pioneer laboratories in 3DEM, and the interrelationships between them.
The genealogy data in the map has purposely been cut off at the year 2011. This arose because of the almost exponential rise in 3DEM activity since this date. We felt that chronicling the early steps of the development of the field would provide a valuable resource in understanding how the field evolved.
Here is a link to the criteria used for inclusion in the genealogy.
We rely on you, our colleagues, to make further contributions to the website and to guide us in the accuracy of the facts we present. Please feel free to contact us (Alexis, Martin, Ardan)!
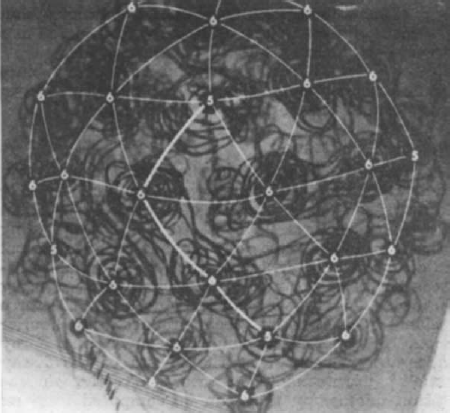
Network Visualization
This is an attempt at an academic genealogy of the field of 3D EM and is a work in progress.
Academic genealogy: Frequently Asked Questions
This genealogy aims to record the growth of the field of 3D EM from 1968 to 2011
Who is included?
Researchers who hold or have held permanent positions and who have made a significant contribution to the field of 3D EM. PhD students, postdocs and other non-permanent scientists are not included. Tenure-track faculty are included.
Technical Staff with more than 5 publications in the field of 3DEM are also included in the list.
What do arrows signify?
Links denote mentorship. Typically, PhD supervisor – student and PI – postdoc relationships are denoted by links. If a person trained or worked in more than one lab, these relationships may be indicated.
Why are some nodes larger, more visible than others?
The choice of which nodes to emphasize aims to reflect:
- Those scientists who initiated the field of 3D EM
- The number of their academic “descendants”
- The fact that some researchers entered the field independently of others, in a sense becoming “first-generation” contributors
Why is X not included? I can see errors, can they be corrected?
Since there is no authoritative source for information needed to compile this genealogy we rely on feedback to ensure there are no omissions or other mistakes.
Development of the field of 3DEM

Publications related to the history of 3D EM
| David DeRosier | 3D reconstruction from electron micrographs a personal account of its development | Methods Enzymol. 2010;481:1-24 |
| Bob Glaeser | Review: Electron Crystallography: Present Excitement, a Nod to the Past, Anticipating the Future | J Struct Biol. 1999 Dec 1;128(1):3-14 |
| Ken Taylor, Bob Glaeser | Retrospective on the early development of cryoelectron microscopy of macromolecules and a prospective on opportunities for the future | J Struct Biol. 2008 Sep;163(3):214-23 |
| Obituary: Walter Hoppe | J. Appl. Cryst. (1987) 20, 324-325 | |
| Bruno Strasser, Jacques Dubochet | Obituary: Eduard Kellenberger (1920-2004) | Nature. 2005 Feb 24;433(7028):817 |
| Marin van Heel | Jean-Pierre Bretaudière (1946-2008) and the early days of multivariate statistics in electron microscopy | In: "An electronic text book: Electron microscopy in Life Science", 3D-EM Network of Excellence, Editors: A. Verkley and E. Orlova (2009) |
| R. Nuzzo | Profile of Chikashi Toyoshima | Proc Natl Acad Sci U S A. 2006 Jan 31;103(5):1165-7 |
| Aaron Klug | Aaron Klug - Autobiography | Nobelprize.org. 17 Jul 2011 |
| Don Caspar, David DeRosier | The 1982 Nobel Prize in chemistry | Science. 1982 Nov 12;218(4573):653-5 |
| John Finch | A Nobel Fellow on Every Floor | Book published by MRC/LMB |
| Anthony Crowther | From Envelopes to Atoms: The Remarkable Progress of Biological Electron Microscopy | Adv Protein Chem Struct Biol. 2010;81:1-32. |
| Viruses and the development of quantitative biological electron microscopy | Notes Rec R Soc Lond. 2004 Jan;58(1):65-81. | |
| Nikolai Andreevich Kiselev | Nikolai Andreevich Kiselev (On the Occasion of His 80th Birthday) | Kristallografiya, 2008, Vol. 53, No. 6, pp. 1149–1150. translated in Crystallography Reports, 2008, Vol. 53, No. 6, pp. 1091–1092 |
| Wolfgang Baumeister | A voyage to the inner space of cells | Protein Sci. 2005 January; 14(1): 257–269. |
| Arthur L Robinson | Electron Microscopy: Imaging Molecules in Three Dimensions | Science 1976 April; Vol. 192 no. 4237 pp. 360-400 |
| Jacques Dubochet | Cryo-EM—the first thirty years | Journal of Microscopy 2011; Vol. 245 no. 3 pp. 1-4 |
| Joachim Frank | Single-particle Cryo-electron Microscopy: The Path Toward Atomic Resolution/Selected Papers Of Joachim Frank With Commentaries (Series in Structural Biology) | April 6, 2018 |
Original personal narratives
These narratives were specially provided to this 3DEM history website by the authors below. We welcome further contributions.
| Robert Josephs | A profile of a researcher in the field of electron crystallography | October 2015 |
| Michael Rossmann | A short scientific autobiography of Michael G. Rossmann | September 2011 |
| Ondreij Krivanek | Ondrej Krivanek’s contribution to microscopy: Memories of an adventure! | August 2018 |
Other Links
Web of stories: video interview of Aaron Klug & Nobel interview with Aaron KlugContributors
Hebrew University of Jerusalem and the National Cancer Institute, NIH
Quick links
Recent Entries
(Show all)2up-1 conformation of HKU1-B S protein after incubation of the receptor
Cryo-EM Structure of CdnG-E2 complex from Serratia marcescens (UltrAuFoil)
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Local map Rrp14/Rrp15/Ssf1 region)
Cryo-EM structure of SV2A dimer in complex with BoNT/A2 Hc and levetiracetam
3up-TM conformation of HKU1-B S protein after incubation of the receptor
Cryo-EM structure of conformation 1 of complex of Nipah virus attachment glycoprotein G with 1E5 neutralizing antibody
Cryo-EM Structure of Spike Glycoprotein from Bat Coronavirus WIV1 in Closed Conformation
Structure of Xenopus tropicalis acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (intermediate state)
60S ribosome biogenesis intermediate (Dbp10 catalytic intermediate - Rrp14/Rrp15/Ssf1 local map)
Closed conformation of HKU1-B S protein after incubation of the receptor
1up-2 conformation of HKU1-B S protein after incubation of the receptor
Xenopus laevis hyaluronan synthase 1, UDP-bound, gating loop inserted state
Chlorella virus Hyaluronan Synthase bound to GlcNAc primer and UDP-GlcA
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (persulfide on NFS1 and ISCU2)
The Anoxybacillus pushchinoensis ORF-less Group IIC Intron HYER1 with 10-nt TRS at symmetric apo state
Xenopus laevis hyaluronan synthase 1, nascent HA polymer bound state
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in the apo-form
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Overall map)
Ensemble map of the Roco protein from C. tepidum in the GTP state bound to the activating Nanobodies NbRoco1 and NbRoco2
CRYO-EM FOCUSED REFINEMENT MAPS OF LEISHMANIA MAJOR 80S RIBOSOME : WILD TYPE REPLICATE 1
Chlorella virus Hyaluronan Synthase bound to GlcA extended GlcNAc primer
Open state of central tail fiber of bacteriophage lambda upon binding to LamB
Roco protein from C. tepidum in the GTP state bound to an activating Nanobody
Cryo-EM Structure of Smooth Muscle Gamma Actin (ACTG2) Mutant R257C
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - Overall map)
Cryo-EM structure of a Legionella effector complexed with actin and AMP
type I-B Cascade bound to a PAM-containing dsDNA target at 3.8 angstrom resolution.
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Local map L1 region)
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology aligning conformation
Mycobacterium tuberculosis ATP synthase F1 in complex with bedaquiline(BDQ)
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology searching conformation
Structure of the dimeric Xenopus tropical acid-sensitive outwardly rectifying channel ASOR trimer bound with tRNA (closed state)
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase F1 in complex with TBAJ-587
Serotonin 1E receptor (5-HT1eR)-Gi1 Complex bound with Setiptiline
Cryo-EM structure of a bacterial nitrilase filament with a covalent adduct derived from benzonitrile hydrolysis
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and brivaracetam
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Low-pass filtered locally refined map)
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (persulfide on ISCU2)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state (full helix)
Cryo-EM structure of a Legionella effector complexed with actin and ATP
1up-1 conformation of HKU1-B S protein after incubation of the receptor
Structure of the human ATP synthase bound to bedaquiline (membrane domain)
Microtubule inner proteins in the 48-nm doublet microtubule from the proximal region of Tetrahymena thermophila strain K40R
Cryo-EM structure of the the 2-oxoglutarate dehydrogenase (E1) with TCAIM complex
Cryo-EM structure of human exon-defined spliceosome in the mature pre-B state.
cryoEM map for design HE0537, a D4 symmetric homo-oligomer designed with RFdiffusion.
Structure of the SARS-CoV-2 EG.5.1 spike glycoprotein in complex with ACE2 (1-up state)
Cryo-EM structure of conformation 2 of complex of Nipah virus attachment G with 1E5 neutralizing antibody
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Peripheral Stalk in complex with TBAJ-587
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - PTC Local map)
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state
Structure of Xenopus tropicalis acid-sensitive outwardly rectifying channel ASOR (resting state)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the second PH domain
Endogenous trans-translation complex with tmRNA*SmpB in the P site and alanyl-tRNA in the A site and deacyl-tRNA in the E site of E. coli 70S ribosome
Monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex in a DNA binding form
Mycobacterium tuberculosis ATP synthase Peripheral Stalk in complex with bedaquiline(BDQ)
Focused map on the Roc-COR domains of the Roco protein from C. tepidum in the GTP state bound to the activating Nanobody NbRoco1
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in complex with TBAJ-587
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase in complex with TBAJ-587
Structural mechanism of inhibition of the Rho transcription termination factor by Rof
Human DNA polymerase theta helicase domain dimer bound to DNA in the microhomology annealed conformation
Cryo-EM structure of human exon-defined spliceosome in the mature B state.
Focused map on the LRR domain of the Roco protein from C. tepidum bound to the activating Nanobody NbRoco2
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and levetiracetam
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state
Cryo-EM structure of human exon-defined spliceosome in the late pre-B state.
Cryo-EM structure of SV2A LD4 in complex with BoNT/A2 Hc in the SV2A-levetiracetam-BoNT/A2 Hc complex
Structure of the human neutral amino acid transporter ASCT2 in complex with nanobody 469
Endogenous trans-translation complex with tmRNA*SmpB in the P site and alanyl-tRNA in the A site of E. coli 70S ribosome
TUBB4B and TUBA1A Heterodimer from Human Respiratory Doublet Microtubules
Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in complex with bedaquiline(BDQ)
Roco protein from C. tepidum in the GTP state bound to the activating Nanobodies NbRoco1 and NbRoco2
Cryo-EM structure of the gasdermin pore from Trichoplax adhaerens
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Dbp10 Local map)
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - L1 local map
Open State of central tail fiber of bacteriophage lambda upon binding to LamB (gpJ713-LamB complex)
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid cyclopentane with a tRNA in pe/E chimeric hybrid state
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid with a tRNA in pe/E chimeric state
Cryo-EM structure of human exon-defined spliceosome in the early B state.
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - Dbp10 Local map)
Cryo-EM Structure of Spike Glycoprotein from Civet Coronavirus 007 in Closed Conformation
Cryo-EM structure of the Pseudomonas aeruginosa PAO1 Type IV pilus
Cryo-EM of the GDP-bound human dynamin (full-length) polymer assembled on the membrane in the super constricted state (full helix)
Cryo-EM of the GDP-bound human dynamin polymer assembled on the membrane in the super constricted state showing the PH domain
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (consensus map)
Structure of the SARS-CoV-2 EG.5.1 spike RBD in complex with ACE2
60S ribosome biogenesis intermediate (Dbp10 pre-catalytic structure - Overall map)
Human DNA polymerase theta helicase domain tetramer in the apo form
Structure of the human mitochondrial iron-sulfur cluster biosynthesis complex during persulfide transfer (without frataxin)
Cryo-EM structure of FLVCR2 in the inward-facing state with choline bound
Cryo-EM structure of SV2A in complex with BoNT/A2 Hc and levetiracetam
Chlorella virus Hyaluronan Synthase bound to GlcA extended GlcNAc primer and UDP
Staphylococcus aureus 70S ribosome with elongation factor G locked with fusidic acid cyclopentane in post-translocational state
2up-TM conformation of HKU1-B S protein after incubation of the receptor
Structure of the human ATP synthase bound to bedaquiline (peripheral stalk domain)
Structure of the human ATP synthase bound to bedaquiline (composite)
Mycobacterium tuberculosis ATP synthase Peripheral Stalk in the apo-form
60S ribosome biogenesis intermediate (Dbp10 post-catalytic structure - H64 Local map)
Cryo-EM structure of FLVCR2 in the outward-facing state with choline bound